Leadered vs. Leaderless Genes: Molecular Mechanisms, Research Methodologies, and Therapeutic Implications
This article provides a comprehensive analysis of the distinctions between leadered and leaderless genes, addressing a critical knowledge gap in prokaryotic and eukaryotic gene regulation.
Leadered vs. Leaderless Genes: Molecular Mechanisms, Research Methodologies, and Therapeutic Implications
Abstract
This article provides a comprehensive analysis of the distinctions between leadered and leaderless genes, addressing a critical knowledge gap in prokaryotic and eukaryotic gene regulation. Tailored for researchers, scientists, and drug development professionals, it synthesizes foundational concepts, advanced detection methodologies, experimental troubleshooting, and comparative validation studies. We explore the unique translation initiation mechanisms, evolutionary significance, and varied genomic prevalence of leaderless genes across species. The content further examines cutting-edge computational and experimental tools for gene structure analysis, addresses challenges in interpreting expression data, and highlights the potential for targeting these distinct genetic structures in therapeutic development, particularly for persistent bacterial infections.
Decoding Genetic Architecture: An Introduction to Leadered and Leaderless Genes
The 5' untranslated region (5' UTR) of messenger RNA (mRNA) represents a critical frontier in the understanding of gene expression regulation. This region, located between the transcription start site and the initiation codon of the main coding sequence, serves as a central hub for post-transcriptional control, influencing mRNA stability, cellular localization, and translation efficiency [1]. The fundamental structural dichotomy in 5' UTR architecture exists between leadered genes, which possess a 5' UTR of varying length and complexity, and leaderless genes, which completely lack this regulatory region and begin directly with the start codon [2]. This distinction represents more than a mere structural curiosity; it embodies divergent evolutionary strategies for translational control with profound implications for basic biological mechanisms and therapeutic development.
Research into leadered versus leaderless genes has revealed that these structures are not randomly distributed across biological kingdoms. While leaderless genes were once considered a rarity in bacteria, genomic analyses have demonstrated they are "totally widespread, although not dominant, in a variety of bacteria," with particularly high proportions in Actinobacteria and Deinococcus-Thermus, where more than twenty percent of genes are leaderless [2]. In archaea, leaderless initiation represents a major mechanism of translation initiation, suggesting deep evolutionary origins [2]. The persistence of this structural dichotomy across domains of life highlights its fundamental importance in gene regulation.
Core Structural and Functional Differences
The architectural distinction between leadered and leaderless genes extends far beyond the simple presence or absence of a 5' UTR, encompassing profound differences in sequence composition, regulatory capacity, and evolutionary trajectory. These differences dictate the very mechanisms by which ribosomes engage with mRNA and initiate the complex process of protein synthesis.
Architectural Composition and Sequence Features
Leadered genes are characterized by a 5' UTR that can vary dramatically in length, from a few nucleotides to several thousand bases [3] [1]. In humans, the median 5' UTR length is approximately 218 nucleotides, representing the longest median length among studied eukaryotes [4]. This expanded regulatory landscape accommodates a complex array of cis-acting elements, including upstream open reading frames (uORFs), upstream AUG codons (uAUGs), secondary structures, and binding sites for proteins and non-coding RNAs [3] [4]. Approximately 42.5% of human 5' UTRs contain at least one uAUG, with 34.4% containing uORFs, 15.0% containing overlapping ORFs (oORFs), and 5.0% containing start-stop elements [3] [5].
In contrast, leaderless genes completely lack these regulatory prefixes, beginning immediately with the initiation codon [2]. This structural minimalism necessitates distinct recognition mechanisms, as the ribosome cannot rely on 5' UTR-mediated guidance to locate the start site. In bacteria, leaderless initiation is often associated with TA-like signals approximately 10-12 base pairs upstream of the translation initiation site, which resemble the -10 box of σ70 factor binding sites and likely represent promoter elements [2].
Table 1: Core Structural Properties of Leadered vs. Leaderless Genes
| Structural Feature | Leadered Genes | Leaderless Genes |
|---|---|---|
| 5' UTR Presence | Present (variable length) | Absent |
| Start Codon Context | Kozak sequence (eukaryotes) or Shine-Dalgarno (some bacteria) | Start codon at transcription start site |
| Regulatory Capacity | High (uORFs, secondary structures, protein binding sites) | Minimal |
| Initiation Mechanism | Scanning (eukaryotes) or direct binding (prokaryotes) | Alternative mechanism, potentially ancient |
| Evolutionary Trend | Increasing complexity in higher eukaryotes | Decreasing proportion in bacterial evolution |
Translational Initiation Mechanisms
The presence or absence of a 5' UTR dictates fundamentally different mechanisms of translation initiation. For leadered genes in eukaryotes, the predominant mechanism is cap-dependent scanning, wherein the 43S pre-initiation complex binds to the 5' cap structure and scans downstream in an ATP-dependent manner until it encounters a suitable start codon [4]. This scanning process can be impeded by RNA secondary structures, which are often unwound by RNA helicases such as eIF4A [4]. The composition of the 5' UTR directly modulates this process; complex secondary structures, uORFs, and specific sequence motifs all influence scanning efficiency and initiation site selection.
Leaderless genes employ a fundamentally different initiation strategy that bypasses many conventional requirements. Experimental evidence suggests that leaderless mRNAs can be faithfully translated across all three domains of life, indicating an ancient and conserved mechanism [2]. This initiation pathway does not require certain canonical initiation factors, and the start codon itself serves as the primary recognition signal. The ribosome appears to bind directly at or near the start codon without extensive 5' end scanning, though the precise molecular details remain an active area of investigation.
Evolutionary Distribution and Trajectory
The distribution of leadered and leaderless genes across the tree of life reveals insightful evolutionary patterns. Comprehensive analysis of 953 bacterial and 72 archaeal genomes demonstrates that leaderless genes are widespread in prokaryotes, though their prevalence varies substantially between lineages [2]. In Actinobacteria and Deinococcus-Thermus, leaderless genes constitute more than 20% of all genes, while in other bacterial groups they appear less frequently.
Evolutionary analysis suggests "that the proportion of leaderless genes in bacteria has a decreasing trend in evolution," indicating that the acquisition of 5' UTRs and the shift toward leadered initiation mechanisms may represent an adaptive refinement of gene regulation [2]. This trend toward increasing regulatory complexity through 5' UTR expansion is particularly evident in eukaryotes, where longer 5' UTRs accommodate more sophisticated regulatory circuits.
Quantitative Analysis of 5' UTR Features and Their Functional Correlates
The structural complexity of 5' UTRs is not random but correlates strongly with functional requirements, particularly for genes requiring precise dosage control. Recent research has quantified these relationships, revealing striking patterns that underscore the regulatory importance of 5' UTR features.
5' UTR Length and Dosage Sensitivity
A comprehensive analysis of 18,764 human 5' UTRs has demonstrated a significant correlation between 5' UTR length and gene dosage sensitivity, as measured by the Loss-of-function Observed/Expected Upper bound Fraction (LOEUF) score [3] [5]. Genes intolerant to loss-of-function (low LOEUF deciles) possess significantly longer 5' UTRs (mean length 269 bp) compared to LoF-tolerant genes (mean length 162 bp; Wilcoxon P<1×10⁻¹⁵) [3] [5]. This relationship remains significant even after controlling for coding sequence length, suggesting that extended 5' UTRs provide expanded regulatory capacity for genes whose expression must be precisely controlled.
Table 2: 5' UTR Features Correlated with Gene Dosage Sensitivity in Human Genes
| 5' UTR Feature | Genes Intolerant to LoF (Low LOEUF) | Genes Tolerant to LoF (High LOEUF) | Statistical Significance |
|---|---|---|---|
| Mean Length | 269 bp | 162 bp | P < 1×10⁻¹⁵ |
| uAUG Content | Higher | Lower | Significant enrichment |
| uORF Content | Higher | Lower | Significant enrichment |
| TSS Diversity | Greater | Less | P < 1×10⁻¹⁵ |
| Secondary Structure Potential | Increased | Reduced | Not specified |
Regulatory Element Composition
The enrichment of regulatory elements in 5' UTRs of dosage-sensitive genes represents a key finding in understanding translational control mechanisms. Upstream AUGs (uAUGs) and upstream open reading frames (uORFs) are significantly enriched in genes intolerant to loss-of-function [3]. These elements generally reduce translation of the downstream main coding sequence, with active uORF translation observed to reduce downstream translation by up to 80% [3] [5]. Ribosome profiling studies have identified that 28.3% of computationally predicted uORFs show evidence of translation, with an additional 45.3% of translated uORFs initiating at non-canonical (non-AUG) start codons [3]. Approximately 20.9% of all 5' UTRs contain one or more ribosome-profiling validated uORFs [3] [5].
The positioning of these regulatory elements within 5' UTRs follows non-random patterns. uORFs are notably depleted in the 100 bp region immediately upstream of the coding sequence, suggesting selective pressure against strongly repressive elements in close proximity to the main start codon [3] [5]. The translation of uORFs themselves is influenced by multiple features, including Kozak sequence strength, local secondary structure, and the distance between the uORF termination codon and the main coding sequence start [3].
Experimental Methodologies for 5' UTR Analysis
Advancements in experimental techniques have been crucial for elucidating the structural and functional properties of 5' UTRs. Both high-throughput screening approaches and mechanistic studies have yielded critical insights into 5' UTR-mediated regulation.
High-Throughput Functional Screening
Modern 5' UTR research has been revolutionized by massively parallel reporter assays (MPRAs) that enable functional characterization of thousands to millions of sequence variants in a single experiment. One sophisticated approach combines polysome profiling of large 5' UTR libraries with deep learning to build predictive models that relate sequence to translation efficiency [6].
In a landmark study, researchers created a library of 280,000 randomized 5' UTRs preceding a constant eGFP coding sequence [6]. After transcribing this library in vitro and transfecting it into HEK293T cells, they separated mRNAs based on ribosome engagement using polysome profiling. By sequencing mRNA from different polysome fractions, they calculated a Mean Ribosome Load (MRL) for each 5' UTR variant, providing a quantitative measure of translation efficiency [6]. This dataset was used to train Optimus 5-Prime, a convolutional neural network that explains 93% of the variance in translation efficiency in held-out test data [6].
To address limitations of lentiviral screening approaches, which are confounded by copy number variations and positional effects, advanced methods employ recombinase-mediated integration strategies that ensure single-copy integration at a defined "landing-pad" location [7]. This approach greatly enhances screening sensitivity by eliminating transcriptional noise, enabling more accurate assessment of 5' UTR regulatory function.
Ribosome Profiling and Translation Initiation Studies
Ribosome profiling (Ribo-seq) has emerged as a transformative method for studying translation at nucleotide resolution. This technique involves nuclease digestion of mRNA not protected by ribosomes, followed by sequencing of the ribosome-protected fragments, thereby providing a genome-wide snapshot of ribosome positions [3] [8].
Application of Ribo-seq has revealed unexpected complexity in 5' UTR translation, including widespread translation of uORFs initiated by both canonical AUG start codons and near-cognate start codons (e.g., CUG, GUG) [3]. In bacteria, ribosome protected footprints exhibit a broad range of lengths (typically 18-40 nucleotides), with the most frequent length being 24 nucleotides in Mycoplasma pneumoniae [8]. These studies have demonstrated that translation initiation rates can vary by over 160-fold among genes in the same organism, highlighting the potent regulatory capacity of 5' UTR sequences [8].
The Scientist's Toolkit: Essential Research Reagents
Table 3: Essential Research Reagents for 5' UTR Investigation
| Reagent / Method | Function in 5' UTR Research | Key Applications |
|---|---|---|
| Polysome Profiling | Separates mRNAs by ribosome number; measures translational efficiency | Genome-wide assessment of ribosome loading [6] |
| Ribosome Profiling (Ribo-seq) | Provides nucleotide-resolution map of ribosome positions | Identifies translated uORFs and initiation sites [3] [8] |
| Massively Parallel Reporter Assays (MPRAs) | Enables high-throughput functional screening of sequence variants | Quantitative analysis of 5' UTR regulatory function [6] |
| Recombinase-Mediated Integration | Ensures single-copy integration at defined genomic sites | Reduces noise in genetic screens [7] |
| Convolutional Neural Networks (CNNs) | Models relationship between sequence and function | Predicts translation efficiency from 5' UTR sequence [6] |
| Hydro-tRNA-seq | Quantifies cellular tRNA abundances | Correlates tRNA availability with translation elongation [8] |
Implications for Therapeutic Development and Synthetic Biology
Understanding 5' UTR structure and function has transcended basic biological significance to become a critical component of therapeutic development and synthetic biology applications. The ability to predict and engineer 5' UTR behavior offers powerful tools for optimizing gene expression in medical and biotechnological contexts.
mRNA Therapeutics and Vaccine Design
In mRNA-based therapeutics, 5' UTR engineering represents a crucial strategy for optimizing protein expression without altering the encoded protein sequence. Research has demonstrated that 5' UTRs can be systematically designed to achieve specified levels of protein production, enabling fine-tuned expression of therapeutic proteins [6]. This approach is particularly valuable for non-viral gene therapies, where maximizing potency is essential for clinical efficacy [7].
Chemical modifications to mRNA, such as pseudouridine (Ψ) or 1-methyl-pseudouridine (m1Ψ), are widely used in therapeutic applications to enhance stability and reduce immunogenicity [6]. Importantly, these modifications can alter the translational properties of 5' UTRs, necessitating specialized models that account for their effects on translation efficiency [6]. The development of predictive models that accommodate modified nucleotides is therefore essential for advancing mRNA therapeutic design.
Variant Interpretation and Disease Pathogenesis
The functional characterization of 5' UTRs has profound implications for understanding human disease. Naturally occurring variants within 5' UTRs can disrupt regulatory elements and cause pathogenic changes in gene expression. Researchers have identified 45 single-nucleotide variants associated with human diseases that substantially alter ribosome loading, suggesting a direct molecular mechanism for pathogenesis [6].
The strong correlation between 5' UTR features and dosage sensitivity provides a framework for prioritizing and interpreting non-coding variants in genetic studies [3] [5]. Genes with complex, regulatory-rich 5' UTRs are more likely to be sensitive to perturbations in this region, highlighting the importance of 5' UTR analysis in diagnostic settings.
Synthetic Biology and Metabolic Engineering
In synthetic biology, 5' UTRs serve as programmable components for fine-tuning gene expression in engineered biological systems. High-throughput screening of synthetic 5' UTR libraries has identified elements that significantly outperform naturally occurring sequences in driving protein expression [7]. These engineered 5' UTRs function across diverse cell types and can be combined to achieve optimal expression levels for specific applications.
In prokaryotic engineering, the distinction between leadered and leaderless architectures provides two distinct strategies for controlling gene expression. Regulatory 5' UTRs that respond to environmental stimuli, such as the ethanol-responsive UTR_ZMO0347 in Zymomonas mobilis, offer dynamic control mechanisms for industrial biotechnology [9]. The modular nature of 5' UTR regulatory elements enables the construction of synthetic genetic circuits that respond to specific metabolic conditions.
The structural dichotomy between leadered and leaderless genes represents a fundamental aspect of gene regulation with far-reaching biological and therapeutic implications. Leadered genes, with their complex 5' UTR architecture, provide an extensive platform for sophisticated regulatory control, particularly for dosage-sensitive genes requiring precise expression. In contrast, leaderless genes employ a minimalist strategy that likely represents an ancient initiation mechanism preserved across evolutionary time.
The investigation of 5' UTR function has been transformed by advanced methodologies including high-throughput screening, ribosome profiling, and machine learning approaches. These techniques have revealed quantitative relationships between sequence features and translational output, enabling predictive models that accelerate both basic research and therapeutic development. As these tools continue to evolve, they will undoubtedly uncover additional layers of complexity in 5' UTR-mediated regulation, further illuminating this critical interface between genetic information and functional proteome.
The initiation of translation, a critical first step in gene expression, occurs through distinct mechanisms in prokaryotes. While the Shine-Dalgarno (SD)-led initiation has long been considered the canonical model in bacteria, leaderless initiation represents a widespread and evolutionarily significant alternative [10] [11]. Leaderless genes are characterized by mRNAs that lack a 5' untranslated region (5' UTR), positioning the translation initiation codon at or very near the 5' end of the transcript [12]. This structural distinction necessitates a different initiation mechanism, where assembled 70S ribosomes, rather than 30S subunits, bind directly to the start codon [11].
Understanding the prevalence and distribution of leaderless genes is not merely a taxonomic exercise. The mechanism is believed to be evolutionarily ancient, potentially used by the last universal common ancestor (LUCA), and is conserved across all three domains of life [10]. Furthermore, because translation initiation is a key regulatory point in gene expression, the leaderless mechanism has profound implications for how gene expression is controlled in pathogens like Mycobacterium tuberculosis and in biotechnologically important organisms like Streptomyces [13] [10]. This guide provides an in-depth technical overview of the distribution of leaderless genes and the experimental methodologies essential for their study, framing this knowledge within the broader context of differentiating leadered and leaderless gene research.
Quantitative Distribution Across Prokaryotes
Large-scale computational analyses have revealed that leaderless genes are not a rarity but a common feature across diverse prokaryotic lineages, though their prevalence varies significantly between archaea and bacteria, and among different bacterial phyla.
Widespread Prevalence in Archaea
In archaea, leaderless initiation is not an alternative but a dominant strategy. Computational studies of multiple complete archaeal genomes indicate that a majority of them possess a substantial proportion of leaderless genes [10]. For instance, transcriptomic studies in species like Pyrobaculum aerophilum and various Haloarchaea have reported that the majority of transcripts are leaderless [10]. This prevalence establishes leaderless initiation as a cornerstone of archaeal gene expression.
Significant Presence in Select Bacterial Phyla
In contrast to archaea, leaderless genes are not dominant in most bacteria, but they are far from uncommon. A comprehensive analysis of 953 bacterial genomes demonstrated that leaderless genes are "totally widespread, although not dominant, in a variety of bacteria" [10]. However, their distribution is highly phylum-specific.
The table below summarizes the prevalence of leaderless genes in key bacterial groups:
Table 1: Prevalence of Leaderless Genes in Select Bacterial Groups
| Bacterial Group / Species | Prevalence of Leaderless Genes | Notes | Source |
|---|---|---|---|
| Actinobacteria (e.g., Mycobacterium, Streptomyces) | >20% of genes | Model organism Streptomyces coelicolor has 18.9% (1,469/7,769 genes) leaderless. [10] | |
| Deinococcus-Thermus | >20% of genes | Noted for a high percentage of leaderless genes. | [10] |
| Mycobacterium tuberculosis | ~14% of annotated genes | Leaderless transcripts are unusually prevalent and translated robustly. | [13] |
| Mycobacterium smegmatis | ~14% of annotated genes | Used as a model organism to study leaderless expression. | [13] |
| Deinococcus deserti | Up to ~60% of genes | Represents an extreme case of leaderless gene abundance. | [11] |
| Escherichia coli | Rare | Commonly known for leadered genes, representing the other end of the spectrum. | [10] [11] |
This phylum-specific distribution suggests an evolutionary trend. Analysis of closely related bacterial genomes implies that the proportion of leaderless genes has a decreasing trend in bacterial evolution, with some lineages retaining a significantly higher fraction than others [10].
Experimental Protocols for Identification and Validation
Accurately identifying leaderless genes and characterizing their expression requires a combination of computational predictions and rigorous experimental validation. Below are detailed methodologies for key experiments in this field.
Computational Identification and Classification
Objective: To genome-widely classify genes as SD-led, leaderless (TA-led), or atypical. Method Summary: This bioinformatic pipeline analyzes sequences upstream of annotated translation initiation sites (TIS).
- Sequence Extraction: Extract 20 bp (for bacteria) or 50 bp (for archaea) of genomic sequence upstream of all annotated TISs [10].
- Signal Detection and Statistical Validation: Use an algorithm designed to detect multi-signals in these upstream regions. The algorithm scans for:
- SD-like signals: Complementary to the 3' end of the 16S rRNA.
- TA-like signals: A "TANNNT" consensus motif resembling a -10 promoter box, typically found ~12 bp upstream of the TIS in bacteria, which indicates a missing 5' UTR and thus a leaderless gene [10].
- The significance of detected signals is validated against shuffled sequences retaining dinucleotide frequency to avoid false positives [10].
- Gene Classification: Classify each gene based on the most probable signal in its upstream sequence:
- SD-led: Possesses a significant SD-like signal.
- TA-led (Leaderless): Possesses a significant TA-like signal at the appropriate distance.
- Atypical: Lacks both significant SD-like and TA-like signals.
Experimental Validation of 5' UTR Boundaries and Translation Initiation
Objective: To empirically determine the transcription start site (TSS) and validate the predicted leaderless structure of a specific gene. Method Summary: This protocol, adapted from studies on M. smegmatis sigA, uses reporter constructs and mutation analysis to confirm the TIS and assess the impact of the 5' UTR [13].
- Reporter Construct Design: Clone the putative promoter and 5' genomic region (including any predicted UTR and the initial part of the coding sequence) of the gene of interest upstream of a fluorescent reporter gene (e.g., Yellow Fluorescent Protein, YFP) in a suitable plasmid vector. The construct is driven by a strong constitutive promoter (e.g., pmyc1tetO) [13].
- Site-Directed Mutagenesis: Systematically mutate putative start codons (e.g., GTG to GTC) within the cloned sequence to identify the true primary translation initiation site [13].
- Expression Analysis: Introduce the reporter constructs (both wild-type and mutated) into the model organism (e.g., M. smegmatis).
- Measurement: Quantify fluorescence (protein abundance), mRNA abundance (via qRT-PCR), and calculate relative transcript production rates.
- Validation: A mutation of the true primary TIS will reduce fluorescence to background levels, confirming its necessity for translation initiation and the leaderless structure if the TIS is at the 5' end [13].
Assessing Translation Efficiency and mRNA Stability
Objective: To directly compare the translation efficiency and mRNA half-life of leadered and leaderless transcripts. Method Summary: This approach uses parallel measurements of protein and mRNA levels over time.
- Strain Construction: Create isogenic strains bearing reporter genes (e.g., YFP) under the control of different 5' UTRs: a leaderless construct, a construct with a long-leadered 5' UTR (e.g., from sigA), and a control with a synthetic 5' UTR [13].
- Transcriptional Arrest: Treat logarithmic-phase cultures with a transcription inhibitor such as rifampin [13].
- Time-Course Sampling: Collect samples at multiple time points post-inhibition.
- Quantitative Analysis:
- mRNA Half-life: Extract total RNA and use qRT-PCR to quantify remaining mRNA levels at each time point. Plot the data to calculate the decay rate and half-life for each construct.
- Translation Efficiency: Measure fluorescence (protein output) and mRNA abundance from untreated cultures. The apparent translation efficiency can be calculated as the ratio of protein abundance to mRNA abundance [13].
- Data Interpretation: Compare the half-lives and translation efficiencies across the different constructs. For example, the sigA 5' UTR may confer a shorter mRNA half-life and decreased apparent translation rate compared to a synthetic leader, while leaderless transcripts may have similar translation efficiency but lower transcript production rates [13].
Research Reagent Solutions
The following table details key reagents and tools essential for experimental research on leaderless genes.
Table 2: Essential Reagents and Tools for Leaderless Gene Research
| Reagent / Tool | Function / Application | Specific Examples / Notes |
|---|---|---|
| Fluorescent Reporter Genes | Quantifying protein abundance and translation efficiency in vivo. | Yellow Fluorescent Protein (YFP) [13]. |
| Constitutive/Inducible Promoters | Driving consistent expression of reporter constructs to isolate post-transcriptional effects. | pmyc1tetO promoter used in mycobacterial systems [13]. |
| qRT-PCR Assays | Measuring absolute and relative mRNA abundance and stability. | Critical for determining mRNA half-life after transcriptional arrest [13]. |
| Transcriptional Inhibitors | Arresting new RNA synthesis to study mRNA decay kinetics. | Rifampin [13]. |
| Bioinformatics Algorithms | Genome-wide prediction and classification of leaderless genes. | Custom algorithms for detecting TA-like signals upstream of TIS [10]. |
| RNA-seq & Ribo-seq | Empirically mapping the 5' ends of transcripts and confirming translation initiation without a 5' UTR. | RNA-seq reads and Ribo-seq reads have coincident 5' boundaries for leaderless genes [12]. |
Visualization of Concepts and Workflows
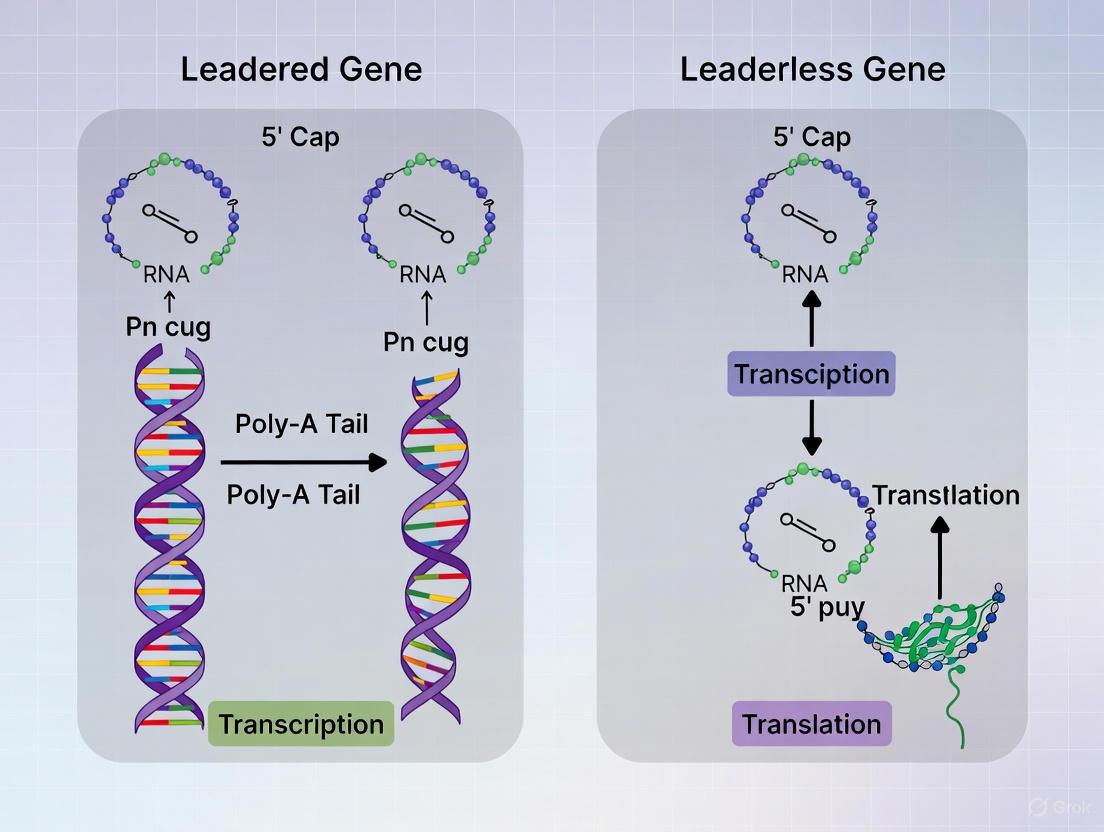
Structural and Mechanistic Differences in Translation Initiation
Diagram 1: Mechanism of leadered versus leaderless translation initiation.
Experimental Workflow for Validating Leaderless Genes
Diagram 2: A multi-step workflow for identifying and validating leaderless genes.
The study of leaderless genes reveals a complex landscape of translation initiation across prokaryotes. Their prevalence, from being widespread in archaea to significant in select bacterial phyla like Actinobacteria, underscores the biological importance of this non-canonical pathway. The distinct mechanism of leaderless initiation, which involves direct 70S ribosome binding and differs in its requirement for initiation factors and specific sequence contexts, represents a fundamental divergence from the SD-led model [10] [11]. For researchers investigating gene regulation, particularly in pathogens like Mycobacterium tuberculosis or industrially relevant organisms like Streptomyces, accounting for leaderless genes is not optional but essential [13] [10]. The experimental and computational frameworks detailed in this guide provide a foundation for exploring this evolutionarily ancient and functionally significant gene class, enabling a more complete understanding of the diversity of life's regulatory strategies.
Leaderless genes, which lack 5' untranslated regions (5'-UTR) and Shine-Dalgarno ribosome-binding sites, represent a molecular relic of ancient translation initiation mechanisms. Once considered a rarity in bacteria, genomic analyses now reveal these genes are widespread across diverse bacterial phyla, though their prevalence shows a marked decreasing trend throughout evolution. This whitepaper examines leaderless genes as molecular fossils within the context of modern gene regulation, highlighting critical differences from leadered genes in translation initiation mechanisms, regulatory constraints, and experimental approaches. We provide quantitative comparisons, detailed experimental protocols for studying both gene types, and essential resources for researchers investigating these ancient genetic elements for drug discovery and synthetic biology applications.
In prokaryotes, translation initiation typically occurs through one of two distinct mechanisms: leadered or leaderless. Leadered genes, which represent the dominant paradigm in well-studied model organisms like Escherichia coli, contain 5'-UTRs with Shine-Dalgarno (SD) sequences that facilitate ribosomal binding through complementary base pairing with the 3'-end of 16S rRNA [10]. In contrast, leaderless genes completely lack 5'-UTRs, with transcription beginning at or immediately adjacent to the start codon, necessitating alternative ribosomal recruitment strategies [14].
The significance of leaderless genes extends beyond their unusual initiation mechanism. Their phylogenetic distribution and structural simplicity suggest they represent an ancient molecular fossil preserved from the earliest stages of cellular evolution. Current evidence indicates that leaderless initiation may be the original translation mechanism used by the last universal common ancestor (LUCA), with the SD-led mechanism representing a more recent evolutionary innovation [10]. This perspective frames the study of leaderless genes not merely as investigation of a biological curiosity, but as a window into primordial gene expression mechanisms.
Evolutionary Significance and Phylogenetic Distribution
Leaderless Genes as Molecular Fossils
The concept of leaderless genes as "molecular fossils" stems from their structural simplicity and universal distribution across all domains of life. The mechanism for translating leaderless mRNAs appears conserved across bacteria, archaea, and eukaryotes, suggesting this capability predates the divergence of these lineages [10]. This conservation, coupled with the minimal requirements for initiation (essentially just a 5'-AUG codon), supports the hypothesis that leaderless translation represents the ancestral state for protein synthesis [14].
Molecular fossils in biology are structures or sequences preserved across evolutionary timescales that provide clues about ancient biological systems. The P-loop found in NTPase proteins represents another example of such a fossil, though its interpretation requires caution as surrounding environmental factors significantly influence its function [15]. Similarly, leaderless genes preserve a simplified translation initiation mechanism that may reflect constraints and opportunities available in early biological systems.
Quantitative Distribution Across Bacterial Phylogeny
Genomic analyses across 953 bacterial genomes reveal that leaderless genes are "widespread, although not dominant, in a variety of bacteria" [10]. However, their distribution is highly uneven across phylogenetic groups:
Table 1: Prevalence of Leaderless Genes Across Bacterial Phyla
| Bacterial Group | Approximate Percentage of Leaderless Genes | Conservation Pattern |
|---|---|---|
| Actinobacteria | >20% | Higher in GC-rich genera |
| Deinococcus-Thermus | >20% | Associated with -10 motif (TANNNT) |
| Other bacterial phyla | Variable (typically <20%) | Generally decreased trend |
| Archaea | Often dominant (>50% in some species) | Ancient, conserved mechanism |
Notably, certain bacterial groups like Actinobacteria (including mycobacteria) and Deinococcus-Thermus exhibit particularly high proportions of leaderless genes, exceeding 20% of their coding sequences [10]. In Mycobacterium tuberculosis and Mycobacterium smegmatis, approximately 14-25% of genes are leaderless [13] [14]. This unusual prevalence in some bacterial lineages suggests either selective pressure maintaining this ancient mechanism or higher rates of leaderless gene formation.
Decreasing Evolutionary Trend
Comparative genomic analyses reveal "the proportion of leaderless genes in bacteria has a decreasing trend in evolution" [10]. This trend is observed when comparing closely related bacterial genomes, where "the change of translation initiation mechanisms... is linearly dependent on the phylogenetic relationship" [10]. The evolutionary trajectory suggests a gradual shift from leaderless-dominated to SD-led initiation mechanisms throughout bacterial evolution, possibly driven by:
- Need for refined regulatory control through 5'-UTRs
- Advantages of ribosomal shielding during translation initiation
- Co-evolution with RNA-based regulatory networks
- Efficiency gains through specialized initiation mechanisms
This decreasing trend parallels the evolutionary fate of many ancient biological systems, which are often supplemented or replaced by more specialized mechanisms while being retained for specific applications where their simplicity provides advantages.
Mechanistic Differences in Translation Initiation
Translation Initiation Pathways
The fundamental distinction between leadered and leaderless genes lies in their translation initiation mechanisms, which employ different ribosomal states, initiation factors, and sequence requirements.
Table 2: Mechanism Comparison Between Leadered and Leaderless Translation
| Characteristic | Leadered Genes | Leaderless Genes |
|---|---|---|
| Ribosomal State | 30S subunit | 70S ribosome (intact) |
| 5'-UTR Requirement | Essential (30-50 nt median) | Absent |
| Key Initiation Factors | IF3, IF1, IF2 | IF2 (enhances), IF3 (inhibits) |
| SD Sequence Role | Critical | Absent |
| Start Codon Position | Internal | 5'-terminal essential |
| Kasugamycin Sensitivity | Sensitive | Resistant [16] |
| mRNA Secondary Structure Sensitivity | High | Low |
The diagram below illustrates the fundamental differences in the translation initiation pathways for leadered and leaderless mRNAs:
Specialized Initiation Mechanisms
Leaderless translation employs specialized mechanisms that distinguish it from canonical initiation:
70S Ribosome Preference: Leaderless mRNAs preferentially bind intact 70S ribosomes rather than 30S subunits, bypassing the subunit association step required for leadered translation [16]. This 70S binding occurs directly at the 5'-terminal start codon without scanning.
Initiation Factor Roles: Initiation factor 2 (IF2/eIF5B ortholog) stimulates leaderless translation, while initiation factor 3 (IF3/eIF3) discriminates against it [17]. This contrasts with leadered initiation, where both factors typically promote efficient initiation.
Alternative Pathways in Eukaryotes: In eukaryotes, leaderless mRNAs can utilize at least four distinct initiation pathways: 80S-mediated, eIF2-dependent, eIF2D-mediated, and eIF5B/IF2-assisted mechanisms [17]. This versatility provides resistance to various cellular stresses that inhibit canonical initiation.
Stress Resistance: Leaderless translation shows relative resistance to certain stressors including kasugamycin antibiotic treatment [16], oxidative stress induced by sodium arsenite, and unfolded protein stress caused by dithiothreitol [17].
Experimental Approaches and Methodologies
Identification and Validation Protocols
Computational Identification Pipeline:
Step 1: Genome Sequence Analysis
- Extract 20-50 bp upstream sequences of all annotated ORFs
- Perform motif discovery using MEME or similar tools [18]
- Identify statistically significant motifs (SD-like: GGAGG; TA-like: TANNNT)
- Classify genes as SD-led, TA-led (leaderless), or atypical
Step 2: Statistical Validation
- Compare identified motifs against shuffled sequences maintaining dinucleotide frequency
- Establish significance thresholds (p < 0.01 typically)
- Validate with known leaderless genes as positive controls
Step 3: Phylogenetic Distribution Analysis
- Map leaderless gene percentages across related genomes
- Correlate with phylogenetic distance
- Identify evolutionary trends in initiation mechanisms
Experimental Validation of Leaderless Transcripts:
Method A: Fluorescence Reporter Assays [13]
- Clone putative 5' regions (including validated start codons) upstream of fluorescent protein genes (YFP/GFP)
- Measure fluorescence intensity as proxy for translation efficiency
- Compare with synthetic control UTRs (e.g., common plasmid UTRs)
- Quantify protein abundance, mRNA abundance, and calculate production rates
Method B: Leaderless Start Codon Verification [13]
- Mutate putative start codons (GTG→GTC, ATG→ATC)
- Measure expression impact via fluorescence or enzymatic activity
- Confirm 5'-terminal position requirement by extending 5' sequence
- Test alternative start codons (ATG, GTG, TTG, ATT) for initiation efficiency
Method C: FLET (FLeeting mRNA Transfection) for Eukaryotic Systems [17]
- Prepare capped, polyadenylated leaderless reporter transcripts (e.g., firefly luciferase)
- Co-transfect with control mRNA (e.g., Renilla luciferase with standard 5'-UTR)
- Measure translation after 2-hour expression window
- Test stress resistance with torin1 (mTOR inhibition), sodium arsenite (oxidative stress), or dithiothreitol (unfolded protein response)
Stability and Expression Measurement Protocols
mRNA Half-Life Determination:
- Treat cultures with transcription inhibitor (rifampin)
- Collect samples at time points (0, 1, 2, 5, 10, 15, 30 minutes)
- Extract RNA, perform quantitative RT-PCR
- Calculate decay rates and half-lives
- Compare leaderless vs. leadered transcript stability [13]
Ribosome Profiling:
- Treat cultures with cycloheximide to arrest ribosomes
- Digest RNA with RNase I (protected fragments ~28-30 nt)
- Purify ribosome-protected fragments
- Construct sequencing libraries
- Map ribosomal positions to transcriptome [14]
Proteomic Validation:
- Perform N-terminal peptide mass spectrometry
- Identify protein N-termini without presumed processing
- Correlate with transcriptional start sites
- Validate small protein expression from leaderless transcripts [14]
Table 3: Essential Research Reagents for Leaderless Gene Studies
| Reagent/Category | Specific Examples | Application/Function | Technical Notes |
|---|---|---|---|
| Reporter Systems | YFP, GFP, Firefly Luciferase | Quantifying translation efficiency | Use promoter-swap constructs to isolate UTR effects |
| Antibiotics | Kasugamycin, Chloramphenicol | Differential inhibition studies | Kasugamycin specifically inhibits 30S but not 70S initiation [16] |
| Stress Inducers | Sodium Arsenite, DTT, Torin1 | Testing translation stress resistance | Arsenite induces eIF2α phosphorylation; DTT causes ER stress |
| Initiation Factors | Recombinant IF2, IF3 | Mechanistic in vitro studies | IF2 enhances leaderless; IF3 inhibits leaderless translation |
| Computational Tools | MEME, RBSfinder, custom algorithms | Identifying leaderless genes in genomes | Look for TANNNT motif ~12 bp upstream in bacteria [18] |
| Model Organisms | M. smegmatis, E. coli, S. cerevisiae | Experimental validation | Mycobacteria have natural high leaderless prevalence (~25%) [14] |
| Ribosome Profiling Kits | Commercial ribo-seq kits | Mapping translating ribosomes | Identifies leaderless ORFs through 5'-terminal ribosome protection |
Research Implications and Applications
Drug Discovery Applications
The unique properties of leaderless translation create promising opportunities for therapeutic intervention:
Selective Antibiotic Targeting: The differential sensitivity of leadered and leaderless translation to antibiotics like kasugamycin suggests potential for pathogen-specific drug development [16]. Compounds could be designed to selectively target the initiation mechanisms predominant in pathogenic bacteria with high leaderless gene content.
Stress Adaptation Targeting: In Mycobacterium tuberculosis, leaderless genes may facilitate adaptation to intracellular stress during infection [13]. Disrupting this adaptive mechanism could enhance host clearance of pathogens.
Small Protein Discovery: Leaderless genes often encode small proteins overlooked by conventional annotation [14]. These represent a largely unexplored repertoire of potential drug targets involved in bacterial physiology and virulence.
Synthetic Biology Applications
Stabilized Expression Systems: Gene fusion strategies that link genes of interest to essential endogenous genes using "leaky" stop codons can enhance evolutionary stability of synthetic constructs [19]. This approach selectively pressures against mutations that disrupt expression.
Regulatory Control: Leaderless architecture simplifies synthetic circuit design by eliminating 5'-UTR regulatory complications. This minimalism facilitates predictable expression in engineered systems.
Heterologous Expression: Understanding leaderless translation mechanisms enables optimization of expression systems for genes from organisms with high leaderless content (e.g., Actinobacteria for antibiotic production).
Leaderless genes represent both a window into evolutionary history and a functionally distinct class of genetic elements with unique regulatory properties. Their decreasing trend throughout evolution marks a transition from ancient, simplified translation mechanisms to more complex, regulated systems. However, their preservation in specific phylogenetic lineages and functional contexts demonstrates ongoing biological relevance.
The structural simplicity of leaderless genes—effectively molecular fossils preserved from early evolution—belies their complex and versatile regulation. Rather than representing imperfect versions of leadered genes, they constitute a parallel system with distinct advantages under specific conditions, particularly stress adaptation. Their study not only illuminates evolutionary history but also reveals alternative biological solutions to fundamental processes like translation initiation.
For researchers and drug development professionals, leaderless genes offer underexplored therapeutic targets and synthetic biology tools. Their differential sensitivity to antibiotics, stress resistance properties, and association with virulence in pathogens present compelling opportunities for intervention. As genomic and proteomic technologies continue advancing, further investigation of these ancient genetic elements will likely yield additional insights with practical applications across biotechnology and medicine.
In the complex machinery of prokaryotic gene expression, the initiation of translation represents a critical control point. This process is fundamentally governed by two distinct paradigms: leadered and leaderless initiation. The Shine-Dalgarno (SD) sequence, discovered by Australian scientists John Shine and Lynn Dalgarno in 1973, is the definitive molecular signature of the canonical leadered pathway [20]. This purine-rich sequence, typically located approximately 8 bases upstream of the start codon AUG, functions as a ribosomal binding site by base-pairing with the complementary anti-Shine-Dalgarno (aSD) sequence at the 3' end of the 16S ribosomal RNA (rRNA) [20] [21]. This interaction precisely aligns the ribosome with the start codon, enabling the formation of the initiation complex and the beginning of protein synthesis.
The recognition that a significant proportion of prokaryotic mRNAs—approximately 14% in mycobacteria and over twenty percent in Actinobacteria and Deinococcus-Thermus—are leaderless (lacking 5'-untranslated regions and SD sequences altogether) has reframed our understanding of translation initiation evolution and mechanisms [13] [2]. This article provides a comprehensive technical examination of the SD-led initiation mechanism, contrasting it with leaderless pathways, and synthesizing current research insights relevant to drug discovery and synthetic biology applications.
Molecular Mechanism of SD-Mediated Initiation
The Core Sequence and Its Recognition
The SD sequence operates through precise molecular complementarity. The consensus six-base sequence is 5'-AGGAGG-3', though variations exist across species and genes [20]. In Escherichia coli, for example, the sequence is typically AGGAGGU, while in T4 phage early genes, the shorter GAGG motif dominates [20]. This sequence base-pairs with the 3'-end of the 16S rRNA, which in E. coli has the pyrimidine-rich sequence 5'-YACCUCCUUA-3' (where Y indicates a pyrimidine) [20] [22].
The effectiveness of this interaction is determined by several key parameters:
- Base-pairing potential: The degree of complementarity between the SD sequence and the aSD sequence directly influences initiation efficiency [22].
- Spacing: The distance between the SD sequence and the start codon is critical, with optimal spacing typically ranging from 5 to 13 nucleotides, and peak efficiency observed at 8-10 nucleotides in E. coli [22].
- Sequence context: Nucleotides surrounding both the SD sequence and start codon contribute to initiation efficiency.
Table 1: Shine-Dalgarno Sequence Variations Across Prokaryotes
| Organism/Group | Core SD Sequence | Anti-SD Sequence on 16S rRNA | Optimal Spacing to Start Codon |
|---|---|---|---|
| Escherichia coli (typical) | AGGAGGU | 5'-AUCACCUCCUUA-3' | 7, 8, 9 bases |
| Bacillus subtilis | GGAGG | 5'-AUCACCUCCUUU-3' | 9, 10, 11 bases |
| T4 phage early genes | GAGG | 5'-AUCACCUCCUUA-3' | ~8 bases |
| Archaea (general) | GGAGG | Shorter variants (e.g., 5'-AUCACCUCC-3') | Variable |
Structural and Functional Dynamics
The primary function of the SD-aSD interaction is to correctly position the ribosome's peptidyl (P) site over the initiation codon, thereby distinguishing the true start codon from internal AUG sequences [23]. This positioning is crucial for translation accuracy. The interaction occurs during the initial stage of 30S ribosomal subunit binding to mRNA, facilitating the subsequent recruitment of initiation factors and the initiator tRNA [20] [21].
The strength of the SD interaction can compensate for other suboptimal features in the translation initiation region. A strong SD sequence can counteract inhibitory mRNA secondary structures that might otherwise block access to the start codon and can also compensate for weak start codons [22]. This compensatory capacity demonstrates the integrative nature of translation initiation control, where multiple sequence elements collectively determine efficiency.
Experimental Analysis of SD Function
Key Methodologies and Reagents
Research into SD-mediated initiation employs a diverse toolkit of molecular, genomic, and computational approaches. The table below outlines essential reagents and methodologies used in contemporary studies.
Table 2: Essential Research Reagents and Methods for Studying Translation Initiation
| Reagent/Method | Function/Application | Key Insights Enabled |
|---|---|---|
| Ribosome Profiling (Ribo-seq) | Genome-wide mapping of ribosome positions | Quantifies translation efficiency across transcriptome; identifies SD-led vs. non-SD initiation [23] |
| Mass Spectrometry (MS) | Detection of novel translated proteins | Identifies proto-genes and unannotated ORFs; validates translation initiation sites [24] |
| Transplastomic Mutants | Introduction of point mutations in aSD sequence | Tests functional relevance of SD-aSD pairing in plastids [23] |
| FLET (Fleeting mRNA Transfection) | Rapid analysis of mRNA translation in living cells | Measures translation efficiency under stress conditions; compares leadered vs. leaderless translation [17] |
| Shuffling Tests with Dinucleotide Frequency | Statistical validation of identified signals | Confirms significance of putative SD sequences above background [2] |
Protocol: Assessing SD Sequence Functionality Through Mutational Analysis
The following protocol outlines a standard approach for experimentally validating SD sequence function:
Step 1: Sequence Analysis and Mutagenesis Design
- Extract the 20-nucleotide region upstream of the start codon from your gene of interest.
- Identify putative SD sequences using motif detection algorithms (e.g., MEME) with statistical validation [2].
- Design mutations that disrupt the SD sequence (e.g., AGGAGG → AGCAGC) while maintaining nucleotide composition similar to wild-type.
- Include compensatory mutations in the 16S rRNA aSD sequence as controls for rescue experiments [20].
Step 2: Reporter Construct Assembly
- Clone the wild-type and mutant 5' UTRs upstream of a reporter gene (e.g., luciferase, GFP) while maintaining the native start codon context.
- Use a low-copy number plasmid with an inducible promoter to control transcription levels.
- Include an internal control (e.g., Renilla luciferase) with a constitutive but distinct 5' UTR for normalization.
Step 3: Transformation and Growth Conditions
- Introduce constructs into an appropriate bacterial strain (e.g., E. coli K-12 derivatives).
- Grow cultures in defined medium to mid-log phase (OD600 ≈ 0.5) under selective pressure.
- Induce expression with sub-saturating inducer concentrations to avoid ribosome limiting conditions.
Step 4: Translation Efficiency Measurement
- Harvest cells and quantify reporter protein levels using enzymatic assays (e.g., luciferase) or immunoblotting.
- Extract total RNA and determine mRNA levels by quantitative RT-PCR to normalize for transcriptional effects.
- Calculate translation efficiency as (reporter protein/mRNA) normalized to internal control.
- For genome-wide studies, employ ribosome profiling to assess ribosomal density at initiation sites [23].
Step 5: Data Interpretation
- Compare translation efficiency between wild-type and SD mutants.
- A significant reduction (typically >50%) confirms SD dependence.
- Test rescue with compensatory aSD mutations in specialized ribosome systems [23].
Genomic Perspectives and Evolutionary Context
Prevalence Across Prokaryotic Lineages
Comparative genomic analyses reveal substantial variation in SD sequence usage across prokaryotic taxa. A comprehensive survey of 30 prokaryotic genomes demonstrated that the presence of SD sequences correlates with multiple gene features, including expression levels, start codon type, and genomic context [22]. The percentage of genes possessing identifiable SD sequences ranges from as low as 10.8% in Mycoplasma genitalium to 90.1% in Thermotoga maritima [22].
This analysis also revealed significant positive correlations between SD presence and predicted expression levels based on codon usage biases. Highly expressed genes are more likely to possess strong SD sequences than average genes, underscoring the importance of efficient initiation for genes whose products are required in large quantities [22]. Additionally, genes with AUG start codons are more likely to possess SD sequences than those with alternative initiators (GUG or UUG), and genes in close proximity to upstream genes show higher SD presence, suggesting operon-specific evolutionary pressures [22].
Evolutionary Relationship to Leaderless Initiation
The evolutionary trajectory of translation initiation mechanisms reveals a fascinating story. Leaderless genes, which completely lack 5' UTRs and therefore SD sequences, are widespread across diverse bacterial lineages, with particularly high abundance in Actinobacteria and Deinococcus-Thermus, where they can exceed 20% of all genes [2]. The proportion of leaderless genes in bacteria shows a decreasing trend in evolution, suggesting that SD-led initiation may represent a more recently derived mechanism that proliferated in specific lineages [2].
The Deinococcus-Thermus phylum exhibits a particularly distinctive expression pattern where a -10 promoter region (TANNNT) is positioned immediately upstream of open reading frames, leading to transcription of leaderless mRNAs without 5' UTRs [18]. This organization suggests an alternative evolutionary pathway where transcription and translation initiation are directly coupled without SD mediation.
Research Applications and Therapeutic Implications
Synthetic Biology and Protein Expression
The predictable nature of SD-aSD interactions makes them invaluable tools for synthetic biology and recombinant protein production. By engineering SD sequences with varying complementarity to the aSD sequence, researchers can precisely tune translation initiation rates to optimize protein expression levels [20]. This principle is extensively applied in bacterial expression systems, where strong SD sequences (e.g., full AGGAGG complementarity) are deployed for high-yield protein production.
The development of orthogonal ribosome systems—where engineered ribosomes with altered aSD sequences specifically translate mRNAs with cognate SD modifications—represents a cutting-edge application of SD mechanics [23]. These systems enable dedicated translation of specific genes independent of cellular regulation, facilitating the production of toxic proteins or the establishment of synthetic genetic circuits.
Antimicrobial Drug Development
The fundamental nature of SD-mediated initiation in bacteria, coupled with its absence in eukaryotic cytoplasmic translation, makes it an attractive target for antimicrobial development. While no approved antibiotics currently target the SD-aSD interaction directly, several promising approaches are under investigation:
- Antisense oligonucleotides designed to block SD sequences on essential bacterial mRNAs
- Small molecules that disrupt the rRNA-mRNA interaction
- Peptide inhibitors that mimic SD sequences and sequester ribosomal binding sites
The taxonomic variation in SD and aSD sequences across bacterial species [22] offers potential for developing narrow-spectrum agents that target specific pathogens while sparing beneficial microbiota. Furthermore, the discovery that leaderless initiation is disproportionately important in certain bacterial taxa (e.g., mycobacteria) suggests that combination therapies targeting multiple initiation mechanisms could overcome resistance [13].
Comparative Integration with Leaderless Initiation
The existence of leaderless mRNAs necessitates a comparative framework for understanding translation initiation. The table below synthesizes key distinctions between these mechanisms.
Table 3: Leadered vs. Leaderless Translation Initiation Mechanisms
| Feature | SD-Led (Leadered) | Leaderless |
|---|---|---|
| 5' UTR | Present (typically 20-50 nt) | Absent or very short |
| SD Sequence | Required for efficient initiation | Absent |
| Ribosome Recruitment | 30S subunit binds via SD-aSD pairing | Can bind 70S ribosomes directly |
| Initiation Factors | IF1, IF2, IF3 in bacteria | Can initiate with IF2 alone or factor-independent |
| Start Codon Context | Spacing from SD critical | First AUG is start codon |
| Evolutionary Prevalence | Dominant in most bacteria | Varies (0.1% to >20% across taxa) [2] |
| Stress Resistance | Standard regulation | Often stress-resistant in eukaryotes [17] |
| Evolutionary Origin | More recent prokaryotic adaptation | Ancient, potentially primordial [2] |
The functional implications of these mechanistic differences are substantial. Leaderless mRNAs demonstrate remarkable resistance to various stress conditions in eukaryotic systems, maintaining translation when canonical initiation factors are compromised [17]. This property may contribute to the persistence of leaderless initiation across evolutionary history despite the proliferation of SD-led mechanisms in many prokaryotic lineages.
The Shine-Dalgarno sequence remains a cornerstone of our understanding of prokaryotic translation initiation, representing the definitive molecular signature of canonical leadered initiation. Its discovery fundamentally shaped molecular biology and continues to inform basic research and applied biotechnology. While the SD mechanism dominates in many bacterial species, the recognition of widespread leaderless initiation across diverse taxa presents a more complex and nuanced picture of translation initiation evolution.
Future research will likely focus on quantifying the dynamic interplay between these initiation mechanisms under varying physiological conditions, mapping the complete network of sequence features that modulate initiation efficiency, and exploiting these fundamental insights for therapeutic development. The continued integration of genomic, biochemical, and structural approaches will further illuminate the intricate molecular ballet that positions ribosomes at the start codon—a process whose precision underpins all cellular life.
Leaderless mRNAs (lmRNAs), which lack 5' untranslated regions (5' UTRs) and Shine-Dalgarno (SD) sequences, represent a significant portion of the transcriptome in diverse organisms, including bacteria, archaea, and eukaryotes. Once considered molecular relics, lmRNAs are now recognized as utilizing sophisticated and diverse translation initiation mechanisms. This whitepaper delineates four distinct translation initiation pathways employed by lmRNAs, a plasticity that contrasts with the more canonical initiation of leadered transcripts. We synthesize current structural, biochemical, and cellular evidence to elaborate the 80S-scanning, eIF2-dependent, eIF2D-mediated, and eIF5B/IF2-assisted pathways. The document provides a comparative analysis of leadered and leaderless initiation, detailed experimental protocols for studying these mechanisms, and a toolkit of essential research reagents. Understanding this mechanistic diversity is paramount for drug development professionals targeting unique translation initiation pathways in pathogens like Mycobacterium tuberculosis, where leaderless genes are exceptionally prevalent.
In the conventional paradigm of prokaryotic gene expression, the 5' untranslated region (5' UTR) of an mRNA plays a critical role in translation initiation by housing the Shine-Dalgarno (SD) sequence, which guides the ribosome to the start codon [13] [25]. Leaderless mRNAs (lmRNAs) defy this paradigm, as they completely lack a 5' UTR and instead possess a start codon at or very near their 5' end. While historically considered rare, genomic and transcriptomic studies have revealed that lmRNAs are widespread across all domains of life.
In bacteria, the prevalence of lmRNAs varies considerably. For instance, in Escherichia coli, they are rare, whereas in organisms like Mycobacterium tuberculosis and Deinococcus deserti, they can represent >20% and up to 60% of all genes [25]. In Mycobacterium smegmatis, approximately 14-25% of genes are leaderless [13] [14]. This abundance in certain bacterial phyla, including Actinobacteria and Deinococcus-Thermus, suggests a significant, non-redundant biological role for leaderless initiation [2]. Archaea and mammalian mitochondria also exhibit a high proportion of lmRNAs, underscoring the evolutionary conservation of this mechanism [17] [26] [2].
The study of lmRNAs is not merely an academic exercise; it is crucial for understanding bacterial adaptation and virulence. M. tuberculosis, a major global pathogen, must alter its gene expression to survive within the hostile environment of its human host [13] [27]. The robust translation of its numerous lmRNAs under stress conditions is a key adaptive strategy. Consequently, the unique translational apparatus required for lmRNAs represents a promising, underexplored target for novel antibacterial therapeutics. This whitepaper explores the four distinct initiation pathways that enable the translation of these unconventional mRNAs.
Four Pathways for Leaderless mRNA Translation Initiation
Eukaryotic systems have evolved multiple strategies to initiate translation on leaderless mRNAs, demonstrating remarkable mechanistic plasticity. These pathways vary in their requirement for initiation factors and the state of the ribosomal subunit, allowing for context-specific regulation.
Table 1: The Four Pathways of Eukaryotic Leaderless mRNA Translation
| Pathway Name | Key Initiating Component | Factor Dependence | Ribosomal State | Key Characteristics |
|---|---|---|---|---|
| 1. 80S-Scanning | Pre-assembled 80S ribosome | eIF2-independent; Resistant to eIF2α phosphorylation | 80S monosome | Initiation factor-free binding; considered an ancient, primordial pathway [17]. |
| 2. eIF2-Dependent | 40S small ribosomal subunit | Requires eIF2, eIF4F, and other canonical factors | 40S subunit | Utilizes the canonical scanning mechanism but on a leaderless template [17]. |
| 3. eIF2D-Mediated | 40S small ribosomal subunit | Dependent on eIF2D, but not eIF2 | 40S subunit | Alternative 48S complex assembly; can function when eIF2 is inactivated [17]. |
| 4. eIF5B/IF2-Assisted | 70S/80S ribosome | Requires eIF5B (eukaryotic homolog of bacterial IF2) | 70S/80S monosome | Mechanistically similar to initiation on certain viral IRES elements; supports initiation under stress [17]. |
The 80S Ribosome-Mediated Pathway
This pathway involves the direct binding of a non-dissociated, intact 80S ribosome to the 5' end of the lmRNA. This mechanism is functionally analogous to the 70S initiation described in bacteria and is remarkably factor-independent. It occurs in the presence of the initiator tRNA, Met-tRNAi, without a requirement for key initiation factors like eIF2 and eIF4F. Consequently, translation via this pathway is highly resistant to cellular stress conditions that inactivate eIF2 (e.g., through phosphorylation by arsenite-induced stress) or impair the eIF4F cap-binding complex [17]. This resilience suggests it serves as an important fail-safe mechanism for maintaining essential protein synthesis during adverse conditions.
The eIF2-Dependent Canonical Pathway
Despite the absence of a 5' leader, lmRNAs can nonetheless engage with the standard cellular translation machinery. In this pathway, the 40S small ribosomal subunit, pre-loaded with initiation factors and the initiator tRNA, binds near the 5' end of the mRNA. It is capable of initiating translation without a scanning process, as the start codon is already positioned at the 5' terminus. This pathway depends on eIF2 for delivering the initiator tRNA and is sensitive to conditions that inactivate this factor [17].
The eIF2D-Mediated Pathway
The eIF2D protein is a non-canonual initiation factor that can deliver the initiator tRNA to the 40S ribosomal subunit independently of eIF2. This provides a third route for lmRNA translation. The eIF2D-mediated pathway becomes particularly important under stress conditions when eIF2 function is compromised, offering an alternative to the canonical eIF2-dependent mechanism [17].
The eIF5B/IF2-Assisted Pathway
The translation initiation factor eIF5B, and its bacterial ortholog IF2, are GTPases that facilitate ribosomal subunit joining. Recent research has uncovered a role for eIF5B in supporting lmRNA translation in eukaryotes. This pathway involves the binding of a 70S/80S ribosome, with the assistance of eIF5B/IF2, and is analogous to the mechanism used by certain viral Internal Ribosome Entry Sites (IRESs), such as that of the Hepatitis C Virus [17]. In bacteria, IF2 is known to stabilize both the initiator tRNA and mRNA binding to the ribosome, and elevated levels of IF2 selectively stimulate lmRNA translation [17] [25].
The coexistence of these four pathways underscores the biological importance of leaderless mRNAs and provides the cell with a versatile regulatory toolkit to fine-tune protein synthesis under diverse physiological conditions.
Structural Insights into Leaderless Initiation
High-resolution structural biology techniques, particularly cryo-electron microscopy (cryo-EM), have provided unprecedented insights into the molecular mechanics of lmRNA translation. A key model system has been the translation of the leaderless λcI mRNA from bacteriophage λ by E. coli ribosomes.
Structural studies of wild-type E. coli 70S ribosomes bound to the λcI lmRNA and initiator fMet-tRNAfMet have confirmed that initiation can occur directly on the intact ribosome without prior subunit dissociation [28] [29]. A critical discovery came from analyzing mutant E. coli strains (e.g., rpsB11) that are deficient in ribosomal protein uS2. These mutants exhibit enhanced translation efficiency of lmRNAs [28] [29].
Cryo-EM structures reveal that uS2-deficient ribosomes also lack ribosomal protein bS21. The absence of these two proteins has profound consequences:
- bS21 normally structurally supports the anti-Shine-Dalgarno (aSD) region of the 16S rRNA. In its absence, the aSD helix is repositioned, which "eases the exit" of the lmRNA from the ribosome's mRNA channel, facilitating stable binding [28] [29].
- The absence of uS2 and bS21 increases the dynamics of the 30S ribosomal head, creating a "peristalsis-like" motion and charge flow within the mRNA entry channel that promotes the propagation of the lmRNA [29].
- A specific π-stacking interaction between the monitor base A1493 of the 16S rRNA and an adenine at the +4 position of the lmRNA may act as a recognition signal for proper start codon positioning [29].
These structural findings explain the long-observed phenomenon of enhanced lmRNA translation in uS2 mutants and highlight the critical role of specific ribosomal proteins in modulating the accessibility of the mRNA exit channel for leaderless transcripts.
Leadered vs. Leaderless Translation: A Comparative Analysis
The fundamental distinction between leadered and leaderless genes—the presence or absence of a 5' UTR—drives profound differences in their expression regulation, from transcription to translation.
Table 2: Comparative Features of Leadered and Leaderless Genes
| Feature | Leadered Genes | Leaderless Genes |
|---|---|---|
| 5' UTR | Present (median ~48-56 nt in mycobacteria) [13] | Absent |
| Shine-Dalgarno (SD) Sequence | Typically present within the 5' UTR | Absent |
| Primary Ribosome Binding Partner | 30S small ribosomal subunit [25] | 70S intact ribosome (in bacteria) [25] |
| Initiation Factor Dependence | High (IF1, IF2, IF3 essential for canonical initiation) [25] | Variable; lower overall. IF3 inhibits; IF2/eIF5B stimulates [17] [25] |
| Start Codon Preference | AUG, GUG, UUG, etc. | Strong preference for AUG (or GTG in some bacteria); strict requirement for 5'-terminal start codon [25] [14] |
| Impact of 5' UTR on mRNA Stability | Significant; secondary structures can protect or destabilize transcripts [13] | Not applicable (no 5' UTR). Stability governed by other features. |
| Transcript Production Rate (in M. smegmatis) | Variable; sigA 5' UTR confers high production rate [13] [27] | Lower predicted transcript production rates [13] [27] |
| Prevalence in M. tuberculosis | ~86% | ~14% - 25% [13] [14] |
Regulatory Consequences for Gene Expression
The structural difference imparts unique regulatory properties to each mRNA type. For leadered transcripts, the 5' UTR is a hub for post-transcriptional regulation. It influences mRNA half-life by forming protective secondary structures or containing motifs that recruit endonucleases [13]. It also modulates translation efficiency via the SD sequence and its accessibility. In mycobacteria, the long 5' UTR of the sigA transcript was shown to cause a short mRNA half-life and decreased apparent translation rate compared to a synthetic control UTR, though it also conferred a higher transcript production rate [13] [27].
In contrast, leaderless transcripts bypass 5' UTR-mediated regulation. Their translation is primarily governed by the affinity of the 70S/80S ribosome for the 5'-terminal start codon and its immediate downstream context. Global studies in M. tuberculosis have found no systematic difference in protein/mRNA ratios between leadered and leaderless transcripts, indicating that variability in translation efficiency is driven by factors beyond leader status [13] [27]. However, their generally lower transcript production rates suggest that transcription initiation is a major point of control for leaderless genes [13].
Experimental Protocols for Investigating lmRNA Translation
The Fleeting mRNA Transfection (FLERT) Assay
The FLERT technique is designed to study translation mechanisms in living mammalian cells while minimizing non-specific effects from transfection and drug treatments [17].
- Reporter Construct Preparation: In vitro transcribe reporter mRNAs encoding a luciferase enzyme (e.g., Firefly luciferase). The test construct is a lmRNA starting with a 5'-AUG-3' (with only one nucleotide prior), while control constructs possess defined 5' UTRs (e.g., human β-actin 5' UTR). All transcripts are capped and polyadenylated.
- Transfection Mix Preparation: Mix the Firefly luciferase test/control mRNAs with a similarly prepared reference mRNA (e.g., Renilla luciferase with a standard 5' UTR) to control for transfection efficiency.
- Cell Transfection: Seed cultured human cells (e.g., HEK293) onto a 24-well plate 12-16 hours before transfection. Transfect the mRNA mixture into the cells using a method that minimally disturbs the culture (e.g., using messengerMAX).
- Stress Induction: Immediately prior to transfection (~5 min), expose cells to various stress conditions to probe pathway dependence:
- eIF2 Inactivation: Add sodium arsenite (e.g., 20-500 µM) to induce oxidative stress and eIF2α phosphorylation.
- eIF4F Inactivation: Treat with Torin1 (e.g., 250 nM) to inhibit mTOR and disrupt the eIF4F complex.
- Unfolded Protein Stress: Treat with dithiothreitol (DTT; e.g., 2 mM).
- Translation Measurement: Harvest cells after a short translation period (e.g., 2 hours). Lyse cells and measure Firefly and Renilla luciferase activities using a dual-luciferase assay system.
- Data Analysis: Normalize Firefly luciferase activity to Renilla activity for each condition. Resistance to stress is indicated by a smaller reduction in normalized lmRNA translation compared to the leadered control mRNA.
In Vitro Reconstitution of Translation Initiation Complexes
This biochemical approach allows for the dissection of specific factor requirements using purified components.
- Ribosome Purification: Purify 70S ribosomes (from bacteria) or 40S/60S subunits and 80S ribosomes (from eukaryotes) from the organism of interest.
- Component Assembly: In a test tube, combine the following in a suitable buffer:
- Ribosomes (70S/80S or 30S/40S).
- Leaderless mRNA transcript (synthesized in vitro).
- Initiator tRNA (fMet-tRNAfMet for bacteria; Met-tRNAi for eukaryotes).
- A defined set of initiation factors (eIF2, eIF5B, eIF2D, etc.)—these are systematically included or omitted to test their necessity.
- GTP (required for factor function).
- Complex Formation: Incubate the mixture at the organism's physiological temperature to allow initiation complex formation.
- Complex Isolation and Analysis:
- Sucrose Density Gradient Centrifugation: Separate formed initiation complexes (30S/48S/70S/80S) from unbound components. Analyze gradient fractions to determine complex size and composition.
- Cryo-EM Grid Preparation: For structural studies, the assembled complex is vitrified on a cryo-EM grid. Data collection and single-particle analysis yield high-resolution structures of the initiation complex, as demonstrated for the λcI lmRNA [28] [29].
Essential Visualizations
Pathway Diagram
The following diagram illustrates the four distinct initiation pathways for leaderless mRNAs, highlighting key differences in ribosomal state and initiation factor requirements.
Experimental Workflow
This flowchart outlines the key steps for the Fleeting mRNA Transfection (FLERT) assay, a key method for studying lmRNA translation in living cells.
The Scientist's Toolkit: Key Research Reagents
Table 3: Essential Reagents for Leaderless mRNA Research
| Reagent / Tool | Function / Utility | Example Use Case |
|---|---|---|
| Reporter Constructs (lmRNA) | Firefly or Nano luciferase transcripts starting with 5'-AUG. Quantifies translation efficiency directly from lmRNA structure. | FLERT assay; in vitro translation efficiency comparisons [17]. |
| Control Reporter Constructs (Leadered) | Luciferase mRNAs with well-defined 5' UTRs (e.g., β-actin). Serves as a benchmark for "standard" translation. | Normalization and stress-resistance calculations in FLERT assays [17]. |
| uS2-Deficient Bacterial Strains | E. coli mutants (e.g., rpsB11) with enhanced lmRNA translation due to altered ribosome structure. | Studying structural requirements and enhanced lmRNA translation mechanisms [28] [29]. |
| Initiation Factor Knockdown/Knockout Systems | siRNA, CRISPR, or inducible knockout systems for factors like eIF2D, eIF5B. | Determining the genetic requirement of specific factors for lmRNA translation in vivo. |
| Specific Inhibitors & Stress Inducers | Sodium Arsenite (induces eIF2α-P), Torin1 (inhibits mTOR/eIF4F), Harringtonine (blocks elongation). | Probing the dependence of lmRNA translation on specific pathways under stress [17]. |
| Cell-Free Translation Systems | Purified, reconstituted systems from bacteria (e.g., E. coli) or eukaryotes (e.g., RRL, HeLa extract). | Biochemical dissection of factor requirements in a controlled environment [17]. |
| Cryo-EM for Structural Biology | High-resolution imaging of ribosome-lmRNA complexes. | Visualizing molecular interactions and conformational changes during lmRNA initiation [28] [29]. |
The study of leaderless mRNAs has moved from the periphery to the forefront of translational control biology. The existence of four distinct initiation pathways—80S-mediated, eIF2-dependent, eIF2D-mediated, and eIF5B/IF2-assisted—underscores the mechanistic diversity and evolutionary importance of this ancient initiation mechanism. This plasticity allows for nuanced regulation of a substantial subset of the genome, particularly in pathogens like Mycobacterium tuberculosis. The structural insights revealing ribosome specialization for lmRNA translation and the development of sophisticated assays like FLERT provide powerful tools for continued discovery. For drug development professionals, the unique molecular machinery required for leaderless initiation, especially the specialized ribosomes and non-canonical factors, presents a promising landscape for developing novel antibacterial agents with new modes of action. Future research will undoubtedly focus on understanding the specific cellular roles of lmRNA-encoded proteins and exploiting this knowledge for therapeutic intervention.
From Sequence to Function: Tools and Techniques for Analyzing Gene Start Features
In the intricate landscape of genomic research, the accurate identification of signals that govern gene expression and protein localization represents a fundamental challenge with profound implications for basic science and drug development. For decades, the scientific community operated under a conventional paradigm where Shine-Dalgarno (SD) sequences and N-terminal signal peptides were considered the dominant regulatory mechanisms in prokaryotes and eukaryotes respectively. However, emerging research has revealed a more complex reality where leaderless genes—those lacking traditional upstream regulatory sequences—are not rare anomalies but widespread features across diverse organisms [10]. This paradigm shift necessitates advanced computational tools capable of detecting these non-canonical genetic signals.
The divergence between leadered and leaderless genes represents more than just a mechanistic curiosity; it touches upon fundamental questions of gene regulation, protein secretion, and evolutionary biology. Leaderless genes, which lack 5'-untranslated regions (5'-UTRs) on their mRNAs, utilize a fundamentally different initiation mechanism where the start codon itself serves as the primary signal for translation [10]. Understanding these differences is critical for multiple applications, including the identification of novel drug targets, optimization of protein expression systems, and reconstruction of evolutionary pathways. This technical guide examines how modern computational algorithms, particularly GeneMarkS-2 and signal peptide predictors, are revolutionizing our capacity to detect and characterize these diverse genetic signals, providing researchers with sophisticated methodologies to navigate this complex terrain.
Biological Background: Leadered versus Leaderless Genes
The Conventional Leadered Gene Paradigm
Traditional understanding of gene architecture in prokaryotes centers on leadered genes characterized by specific upstream regulatory elements:
- Shine-Dalgarno (SD) sequences: Short motifs in the 5'-UTR that base-pair with the 3'-end of 16S rRNA to facilitate ribosome binding and translation initiation [10]
- 5'-untranslated regions (5'-UTRs): Variable-length sequences preceding the start codon that contain regulatory information
- Promoter elements: Including -10 Pribnow box (TATAAT) and -35 elements in bacteria that direct transcription initiation
This leadered architecture has dominated microbiological textbooks and computational models for decades, with most gene prediction algorithms heavily relying on these features for accurate gene calling.
The Emerging Leaderless Gene Paradigm
In contrast, leaderless genes challenge this conventional model through their distinctive characteristics:
- Absence of 5'-UTRs: The transcription start site coincides with the first nucleotide of the translation initiation codon [10]
- Lack of SD sequences: Translation initiation occurs without ribosome binding to upstream SD motifs [10]
- Simplified initiation mechanism: The start codon itself serves as the primary recognition signal for translation machinery [10]
- Evolutionary conservation: Evidence suggests this initiation mechanism may be used by the last universal common ancestor (LUCA) and conserved across all domains of life [10]
Research has demonstrated that leaderless genes are not merely rare exceptions but constitute significant proportions of genomic content across diverse taxa. In Actinobacteria and Deinococcus-Thermus, for instance, over twenty percent of genes are leaderless [10]. This prevalence underscores the biological significance of this initiation mechanism and the necessity of computational tools capable of detecting these genes.
Computational Tools and Algorithms
GeneMarkS-2: Advanced Prokaryotic Gene Prediction
GeneMarkS-2 represents a significant advancement in ab initio gene prediction for prokaryotic genomes. Unlike earlier tools that primarily relied on canonical SD-led initiation signals, GeneMarkS-2 incorporates multiple sequence patterns including those characteristic of leaderless transcription [30] [31]. The algorithm employs a sophisticated self-training approach to derive species-specific (native) models while utilizing precomputed heuristic models to identify harder-to-detect genes, including those likely to have been horizontally transferred [30].
Key innovations of GeneMarkS-2 include:
- Dual-model system: Native models for species-specific genes and heuristic models for horizontally transferred genes
- Leaderless transcription modeling: Explicit incorporation of non-canonical initiation patterns
- Noncanonical RBS recognition: Ability to identify ribosome binding sites that deviate from SD consensus
- Comprehensive signal detection: Identification of various sequence motifs regulating gene expression
Benchmarking tests have demonstrated that GeneMarkS-2 outperforms previous state-of-the-art tools in all accuracy measures when validated against genes confirmed by COG annotation, proteomics experiments, and N-terminal protein sequencing [30].
SignalP 6.0: Comprehensive Signal Peptide Prediction
For protein secretion signal detection, SignalP 6.0 represents the current gold standard. This tool utilizes a transformer protein language model with a conditional random field for structured prediction, enabling identification of all five known types of signal peptides across all domains of life [32] [33].
The key advancements in SignalP 6.0 include:
- Expanded classification capability: Detection of Sec/SPI, Sec/SPII, Sec/SPIII, Tat/SPI, and Tat/SPII signal peptides [33]
- Protein language models: Leveraging contextual representations from millions of unannotated protein sequences [33]
- Region identification: Precise delineation of n-region, h-region, and c-region within signal peptides [33]
- Organism-agnostic prediction: Accurate performance even without prior knowledge of species origin [33]
SignalP 6.0 shows particular improvement in detecting SP types with limited training data (Sec/SPIII and Tat/SPII) and generalizes better to evolutionarily distant proteins compared to previous versions [33].
Complementary Tool Landscape
Table 1: Computational Tools for Genetic Signal Detection
| Tool Name | Primary Function | Key Features | Organism Scope |
|---|---|---|---|
| GeneMarkS-2 | Gene prediction | Leaderless transcription modeling, self-training, heuristic models | Prokaryotes (Bacteria, Archaea) |
| SignalP 6.0 | Signal peptide prediction | Five SP-type classification, protein language models, region identification | All domains of life |
| DeepLoc/DeepLocPro | Subcellular localization | Localization prediction, complementary to SP prediction | Eukaryotes/Prokaryotes |
Experimental Protocols and Methodologies
High-Throughput Signal Peptide Efficiency Screening
Recent advances have enabled large-scale experimental characterization of signal peptide functionality through innovative screening methodologies:
- Library Design: Rational design of 11,643 unique SP variants based on 134 wild-type SPs from B. subtilis, modifying physicochemical features while minimizing effects on correlated properties [34]
- Nanoliter Reactor (NLR) Screening: Encapsulation of library strains in NLRs for high-throughput quantification of secretion efficiency via amylolytic degradation of fluorescein-labeled starch [34]
- Machine Learning Integration: Training of random forest models using 156 informative physicochemical features to predict secretion efficiency [34]
- Feature Importance Analysis: Application of TreeSHAP for post-hoc explanation of model predictions and identification of critical SP features [34]
This integrated approach demonstrated superior sensitivity compared to traditional microtiter plate assays and enabled correlation of specific physicochemical features with secretion efficiency.
Computational Identification of Leaderless Genes
The detection of leaderless genes in bacterial genomes employs specialized computational workflows:
- Sequence Extraction: Retrieval of 20 bp translation initiation site (TIS) upstream sequences for all protein-coding genes in a genome [10]
- Multi-signal Classification: Algorithmic classification of genes into SD-led, TA-led, and atypical categories based on predominant upstream signals [10]
- Statistical Validation: Shuffling tests to confirm statistical significance of identified signals compared to dinucleotide-frequency preserved random sequences [10]
- Genome-Wide Annotation: Application across 953 bacterial and 72 archaeal genomes to identify leaderless genes based on TA-like signals approximately 10 bp upstream of TIS [10]
This methodology revealed that TA-like signals in bacteria (consensus TANNNT) resemble the -10 box of σ70 factor binding sites and indicate very short or missing 5'-UTRs, characteristic of leaderless genes [10].
Experimental Validation of Signal Peptide Predictions
Rigorous experimental validation remains essential for benchmarking computational predictions:
- Empirical Determination: Large-scale N-terminal sequencing of 270 secreted recombinant human proteins via automated Edman analysis [35]
- Algorithm Benchmarking: Evaluation of prediction tools (SignalP, SigCleave, SigPfam) against experimentally verified cleavage sites [35]
- Database Annotation Assessment: Comparison of computational predictions with SWISS-PROT annotations to identify discrepancies [35]
- Model Refinement: Use of verified data to improve prediction accuracy through refined profile hidden Markov models [35]
These validation efforts revealed that only 70.5% of annotated cleavage sites in SWISS-PROT agreed with experimental data, with the agreement rate rising to 85.0% for entries specifically marked as experimentally verified rather than computationally predicted [35].
Data Analysis and Visualization
Quantitative Analysis of Tool Performance
Table 2: Performance Comparison of Signal Peptide Prediction Tools
| Tool Version | Cleavage Site Accuracy | Key Improvements | Limitations |
|---|---|---|---|
| SignalP 2.0-NN | 78.1% [35] | Neural network method | Limited SP type discrimination |
| SignalP 4.0 | Not quantified | Better discrimination from transmembrane regions | Unable to detect all SP types |
| SignalP 5.0 | Not quantified | Discrimination of 3 SP types using deep neural networks | Cannot detect Sec/SPIII or Tat/SPII |
| SignalP 6.0 | Substantial precision gains [33] | All 5 SP types; better generalization | Relies on correct start codon identification |
Evolutionary Distribution of Leaderless Genes
Analysis of 953 bacterial genomes reveals striking taxonomic patterns in leaderless gene distribution:
- Actinobacteria and Deinococcus-Thermus: Over 20% of genes are leaderless [10]
- Streptomyces coelicolor A3(2): 18.9% (1469 of 7769 genes) classified as leaderless [10]
- Evolutionary Trend: Decreasing proportion of leaderless genes throughout bacterial evolution [10]
- Functional Correlation: Translation initiation mechanism changes correlate with phylogenetic relationships [10]
These distribution patterns suggest evolutionary trade-offs between different initiation mechanisms and potential adaptive significance in specific environmental contexts.
Table 3: Key Experimental Reagents and Computational Resources
| Resource Name | Type | Function/Application | Access Information |
|---|---|---|---|
| GeneMarkS-2 Web Server | Computational tool | Prokaryotic gene prediction with leaderless transcription modeling | [31] |
| SignalP 6.0 Server | Computational tool | Multi-type signal peptide prediction | [32] |
| NLR Assay System | Experimental platform | High-throughput secretion efficiency screening | [34] |
| SP Library (11,643 variants) | Research reagent | Benchmarking SP sequence-function relationships | [34] |
| Signal Peptide Training Datasets | Data resource | Model training and benchmarking | [36] [32] |
Technical Implementation and Workflow
Integrated Computational-Experimental Pipeline
The following diagram illustrates a comprehensive workflow for genetic signal detection that integrates computational prediction with experimental validation:
Integrated Workflow for Genetic Signal Detection
Signal Peptide Structure and Detection
The structural features of signal peptides and their computational detection are visualized in the following diagram:
Signal Peptide Structure and Detection Methodology
The integration of advanced computational algorithms like GeneMarkS-2 and SignalP 6.0 represents a transformative development in the detection and characterization of genetic signals. These tools have moved the field beyond the simplistic leadered-gene paradigm to embrace the complexity and diversity of genomic regulation in all domains of life. The ability to accurately identify leaderless genes and diverse signal peptide types has profound implications for understanding fundamental biological processes, including gene regulation, protein secretion, and evolutionary mechanisms.
For drug development professionals, these advances offer new opportunities for target identification, particularly for secreted proteins and membrane-associated receptors that represent prime therapeutic targets. The improved accuracy in predicting signal peptides facilitates the identification of surface proteins in pathogens, potentially revealing novel antibiotic targets. Furthermore, the capacity to optimize signal peptides for recombinant protein production holds significant promise for biopharmaceutical manufacturing.
Future developments will likely focus on the integration of multi-omics data, improved prediction of condition-specific gene expression, and application to metagenomic datasets of unknown origin. As protein language models continue to evolve, we can anticipate further improvements in detection accuracy, particularly for rare signal peptide types and across evolutionary distant sequences. The continuing dialogue between computational prediction and experimental validation will remain essential for refining these tools and expanding their applications in basic research and drug development.
The classical model of prokaryotic translation initiation, dominated by the Shine-Dalgarno (SD) mechanism, has been fundamentally redefined by the recognition of alternative pathways. For decades, the SD sequence was considered the universal bacterial mechanism for ribosome binding, facilitating start codon recognition through base-pairing with the 3'-end of the 16S rRNA [2]. However, genomic analyses have revealed that non-SD-led genes are as common as SD-led genes across prokaryotes [37], demanding systematic approaches to classify initiation mechanisms. Research now distinguishes between "leadered" genes, which possess 5'-untranslated regions (5'-UTRs) containing regulatory signals, and "leaderless" genes, which lack 5'-UTRs and thus initiate translation through fundamentally different mechanisms [2]. This classification is not merely academic; it provides crucial insights into evolutionary conservation, with leaderless initiation potentially representing the ancestral mechanism used by the last universal common ancestor (LUCA) and conserved across all domains of life [2]. Accurately annotating these initiation patterns is therefore essential for understanding gene regulation, optimizing heterologous protein expression, and identifying novel drug targets in pathogenic bacteria.
Core Principles: Defining Initiation Archetypes
Prokaryotic translation initiation mechanisms fall into three primary categories, each defined by distinct sequence signatures upstream of the translation initiation site (TIS).
SD-led (Leadered) Genes: These genes contain a 5'-UTR with a Shine-Dalgarno sequence, typically the consensus GGAGG or AGGAGG, located 5-10 nucleotides upstream of the start codon [18]. This motif base-pairs with the anti-SD sequence at the 3'-end of the 16S rRNA, positioning the ribosome for accurate initiation [38]. This mechanism has long been considered dominant in bacteria and allows for tunable translation rates based on SD::aSD binding strength.
TA-led (Leaderless) Genes: These genes lack a 5'-UTR and are instead characterized by a TA-rich motif, often with the consensus TANNNT, positioned approximately 10-12 bp upstream of the TIS in bacteria [2] [18]. This motif closely resembles the -10 box (Pribnow box) of σ70-dependent bacterial promoters [2]. For these genes, the TA-like signal functions primarily as a transcriptional promoter element, with transcription initiating at or very near the start codon, resulting in leaderless mRNA [18]. Translation initiation then occurs directly at the 5'-end of the mRNA without the need for ribosome recruitment via SD pairing.
Atypical Genes: A significant proportion of genes do not contain strong SD or TA signals in their upstream regions [2]. These atypical genes may utilize non-canonical initiation mechanisms, which could include RPS1-mediated initiation, translational scanning, or internal ribosome entry sites (IRES), mechanisms more traditionally associated with eukaryotes [38]. The precise signals governing their initiation remain an active area of research.
Table 1: Core Characteristics of Translation Initiation Types
| Initiation Type | Key Upstream Signal | 5'-UTR Status | Primary Functional Role | Prevalence Example |
|---|---|---|---|---|
| SD-led (Leadered) | Shine-Dalgarno (e.g., GGAGG) | Present | Translational (Ribosome Binding) | ~90% in Bacillus subtilis [38] |
| TA-led (Leaderless) | TA-like motif (e.g., TANNNT) | Absent or very short | Transcriptional (Promoter) | >20% in Actinobacteria [2] |
| Atypical | Weak or no discernible SD/TA signal | Variable | Unknown/Alternative Mechanisms | ~50% in Caulobacter crescentus [38] |
Computational Methodologies for Signal Annotation
Large-scale identification and classification of translation initiation signals require robust bioinformatic pipelines that combine motif discovery, statistical validation, and comparative genomics.
Signal Detection and Classification Algorithms
A powerful approach involves a MEME-like algorithm that integrates several parameters into a likelihood function for signal identification [37]. This method combines:
- Position Weight Matrix (PWM) of the potential signal.
- Spacer length distribution between the signal and the TIS.
- Genomic background nucleotide frequencies.
An Expectation-Maximization (EM) algorithm and simulated annealing are used for parameter estimation [37]. Classification is then performed by scoring each detected signal against reference PWMs for SD consensus ("AAGGAGGTGA") and the Pribnow box (from E. coli K-12) or a TATA box (from archaea). Signals are categorized based on their resemblance to these references [37].
Statistical Validation and Phylogenetic Analysis
To ensure biological significance rather than random occurrence, signals must be validated against null models. This is achieved by comparing observed signal strength (e.g., the fraction of TA-led genes, f~TA,obs~) to an expected value derived from hundreds of nucleotide-shuffled genomes (f~TA,rand~) [2]. The statistically significant signal is then quantified as Δf~TA~ = f~TA,obs~ - f~TA,rand~ [2].
For broader evolutionary insights, sequence entropy analysis can be applied. The information content (ΔI) across the initiation region (e.g., positions -20 to -4 relative to the TIS) summarizes position-specific conservation and effectively quantifies genome-wide SD sequence utilization, correcting for uneven nucleotide usage [38].
Figure 1: Computational Workflow for Initiation Signal Annotation. The pipeline begins with genomic sequence input and progresses through motif discovery and statistical validation to final classification.
Experimental Validation Protocols
Computational predictions require rigorous experimental validation. The following protocols are essential for confirming the function of predicted SD-led and TA-led initiation signals.
Reporter System Construction and Mutagenesis
To empirically validate a predicted TA-led promoter, a standard approach involves cloning the upstream region containing the TANNNT motif directly in front of a promoterless reporter gene (e.g., GFP or lacZ) without introducing a Shine-Dalgarno sequence [18]. The critical control is a construct with site-directed mutations in the conserved residues of the -10 motif (e.g., TANNNT → GCNNNT) [18]. A significant reduction in reporter expression in the mutated construct confirms the functional importance of the motif. Furthermore, to test if the sequence can drive transcription of leaderless mRNA, 5'-RACE (Rapid Amplification of cDNA Ends) can be performed to identify the transcriptional start site (TSS). A TSS coinciding with the first nucleotide of the start codon provides definitive evidence for a leaderless gene architecture [2] [18].
Proteomic Validation of Initiation Sites
For direct, genome-scale experimental validation of TIS, mass spectrometry (MS) is the gold standard. Peptides detected via MS can confirm the N-terminal amino acid sequence of a protein, thus validating the predicted start codon [24]. Key considerations for this approach include:
- Using high-confidence thresholds (e.g., q-value < 0.0001) to minimize false positives from non-coding ORFs [24].
- Manual curation of fragmentation spectra is often required for novel or short proteins [24].
- Integration with ribosome profiling (Ribo-seq) data can provide additional evidence of translation, though it is more prone to artifacts from stochastic expression [24].
Table 2: Essential Reagents and Resources for Experimental Validation
| Research Reagent / Method | Critical Function | Application Context |
|---|---|---|
| Promoterless Reporter Vector (e.g., GFP, lacZ) | Provides a measurable readout for promoter and initiation region activity. | Functional validation of predicted TA-led promoters [18]. |
| Site-Directed Mutagenesis Kit | Introduces precise mutations (e.g., TANNNT → GCNNNT) in putative motifs. | Determining the necessity of conserved motif residues [18]. |
| 5'-RACE (Rapid Amplification of cDNA Ends) | Precisely maps the Transcriptional Start Site (TSS). | Confirming leaderless transcription (TSS = start codon) [2] [18]. |
| High-Sensitivity Mass Spectrometry | Detects and sequences translated peptides from cellular extracts. | Experimental verification of Translation Initiation Sites (TIS) at proteome scale [24] [37]. |
| Ribosome Profiling (Ribo-seq) | Captures ribosome-protected mRNA fragments, indicating active translation. | Genome-wide identification of translated ORFs; can complement MS data [24]. |
Genomic Distribution and Evolutionary Trends
Large-scale surveys across hundreds of prokaryotic genomes reveal that leaderless genes are a widespread phenomenon, though their prevalence varies dramatically across phylogenetic groups.
Taxonomic Variation in Initiation Mechanisms
Analysis of 953 bacterial and 72 archaeal genomes demonstrates that while SD-led initiation is dominant in many model organisms, leaderless genes are abundant in specific phyla [2]. For instance, over twenty percent of genes are leaderless in Actinobacteria and Deinococcus-Thermus [2]. Within the Deinococcus-Thermus phylum, the -10 motif (TANNNT) adjacent to the ORF represents a common expression pattern, responsible for transcribing a significant proportion of leaderless genes [18]. In contrast, some bacterial species like Caulobacter crescentus possess a high fraction (∼50%) of genes that are non-SD-led, many of which fall into the atypical category [38].
Correlates and Evolutionary Drivers
The propensity for different initiation mechanisms correlates with several genomic and lifestyle traits. Species with fast growth rates and high protein production demands tend to have a greater proportion of SD-led genes, likely because the SD mechanism allows for finer tuning of translation initiation rates to optimize efficiency [38]. Furthermore, environmental factors play a role; thermophilic species contain significantly more SD-led genes than mesophiles, potentially because stronger SD::aSD pairing stabilizes ribosome-mRNA interactions at high temperatures [38]. Macroevolutionary analysis suggests that the proportion of leaderless genes in bacteria has followed a decreasing trend throughout evolution [2], with SD-led initiation potentially being a more recent adaptation in many lineages.
Table 3: Prevalence of Leaderless Genes Across Prokaryotic Groups
| Taxonomic Group | Approximate Proportion of Leaderless Genes | Key Genomic/Environmental Correlates |
|---|---|---|
| Actinobacteria | >20% [2] | --- |
| Deinococcus-Thermus | >20% [2] | Extraordinary environmental adaptability [18]. |
| Escherichia coli | Minor fraction [2] | Fast growth rate, model lab organism. |
| Thermophiles | Lower (Higher SD-led proportion) [38] | High optimal growth temperature. |
| Archaea | Highly abundant, often dominant [2] | --- |
Implications for Genome Annotation and Synthetic Biology
The existence of multiple initiation mechanisms has profound practical implications for both basic and applied microbiology.
Accurate genome annotation is critically dependent on correctly identifying TIS. Misannotation can occur if a leaderless gene is predicted to have a long 5'-UTR based on an upstream SD-like sequence that is actually a promoter element for a downstream gene within an operon [2]. Specialized databases like ProTISA have been developed to catalog confirmed TISs using a combination of experimental data, conserved domain analysis, and homology mapping, providing a refined resource beyond standard automated annotation [37]. In synthetic biology, understanding these natural variations is key to engineering robust gene expression systems. For organisms like Deinococcus radiodurans, synthetic constructs can be designed leveraging the minimal -10 motif for basic expression, with the addition of a -35 region to significantly enhance transcriptional output [18]. Similarly, optimizing the strength and spacing of the SD sequence is a established strategy for tuning protein production in traditional bacterial chassis like E. coli [38].
In the field of genomics, establishing statistical significance is paramount for distinguishing authentic biological signals from random noise. Shuffling tests, also known as permutation tests, provide a robust computational framework for this purpose, offering a non-parametric approach to hypothesis testing that does not rely on strict distributional assumptions. These methods are particularly valuable in "leadered versus leaderless" genes research, where researchers investigate fundamental differences in gene regulation mechanisms between transcripts containing 5' untranslated regions (5'-UTRs) and those that lack them entirely.
The core principle of shuffling tests involves systematically randomizing observed data to create an empirical distribution of a test statistic under the null hypothesis. In the context of signal detection for gene research, this typically involves randomizing sequence positions or labels to determine whether an observed signal—such as an overrepresented motif in a genomic region—occurs more frequently than would be expected by chance alone. This approach is especially useful for validating putative regulatory signals identified in genomic analyses, where traditional parametric tests may be inappropriate due to unknown sampling distributions or complex dependencies within the data.
Within leadered and leaderless gene research, shuffling tests have enabled scientists to address critical questions about the evolutionary conservation of translation initiation mechanisms, the statistical significance of identified promoter motifs, and the functional implications of different gene structures. As research in this field progresses, employing rigorous statistical validation methods like shuffling tests becomes increasingly important for generating reliable, reproducible findings that advance our understanding of gene regulation in prokaryotes and beyond.
Biological Context: Leadered vs. Leaderless Genes
Fundamental Differences in Translation Initiation
In prokaryotic systems, genes are primarily categorized into two classes based on their translation initiation mechanisms: leadered and leaderless genes. Leadered genes contain 5' untranslated regions (5'-UTRs) that typically include a Shine-Dalgarno (SD) sequence, which facilitates ribosome binding and translation initiation through base-pairing with the 3'-end of the 16S rRNA [13]. In contrast, leaderless genes completely lack 5'-UTRs, with the transcription start site located at or immediately adjacent to the translation initiation codon [2]. This structural difference necessitates distinct translation initiation mechanisms, as leaderless genes cannot rely on SD-mediated ribosome binding.
While SD-led initiation has long been considered the dominant translation mechanism in prokaryotes, genomic analyses have revealed that leaderless genes are surprisingly widespread across bacterial taxa. Approximately 14% of annotated genes in both Mycobacterium smegmatis and Mycobacterium tuberculosis are leaderless, with some bacterial phyla like Actinobacteria and Deinococcus-Therpus exhibiting even higher proportions exceeding 20% [13] [2]. This prevalence suggests that leaderless translation represents a functionally important alternative initiation mechanism rather than a rare exception.
Functional and Regulatory Implications
The structural differences between leadered and leaderless genes have profound implications for their regulation and functional properties. Leadered transcripts with extended 5'-UTRs can accommodate complex regulatory elements, including binding sites for small RNAs, RNA-binding proteins, and riboswitches that modulate translation efficiency and mRNA stability [13]. Research has demonstrated that 5'-UTRs can significantly impact transcript stability, with the sigA 5'-UTR in mycobacteria conferring decreased mRNA half-life compared to synthetic controls [13].
Leaderless genes, lacking these extended regulatory regions, appear to employ fundamentally different regulatory strategies. Evidence suggests that leaderless transcripts in mycobacteria are translated with similar efficiency as their leadered counterparts, though they may be transcribed less efficiently, resulting in lower steady-state mRNA and protein abundances [13]. The absence of 5'-UTRs also has implications for transcription-translation coupling, a fundamental feature of prokaryotic gene expression where the processes of transcription and translation are physically and functionally linked.
Table: Comparative Features of Leadered and Leaderless Genes
| Feature | Leadered Genes | Leaderless Genes |
|---|---|---|
| 5'-UTR Presence | Present (median 48-56 nt in mycobacteria) | Absent |
| SD Sequence | Typically present | Absent |
| Translation Initiation Mechanism | SD-mediated ribosome binding | Start codon recognition |
| Representation in Mycobacteria | ~86% of genes | ~14% of genes |
| Regulatory Capacity | Complex (sRNAs, protein binding) | Limited |
| mRNA Stability | Variable (influenced by 5'-UTR) | Generally stable |
| Evolutionary Trend | Increasing | Decreasing |
Statistical Foundations of Shuffling Tests
Conceptual Framework and Algorithmic Implementation
Shuffling tests belong to the family of permutation tests, which operate on the fundamental principle of randomly rearranging observed data to create an empirical distribution of a test statistic under the null hypothesis. The core algorithm involves: (1) calculating the test statistic for the observed data, (2) repeatedly shuffling the data labels or values to create simulated datasets under the null hypothesis, (3) recalculating the test statistic for each shuffled dataset, and (4) determining the proportion of shuffled datasets that produce test statistics as extreme as or more extreme than the observed statistic [39]. This proportion constitutes the empirical p-value, providing a direct measure of statistical significance without relying on parametric assumptions.
The implementation of shuffling tests requires careful consideration of the appropriate null model and shuffling unit. In genomic applications, common approaches include sequence shuffling that preserves local nucleotide composition while randomizing higher-order patterns, or label shuffling that randomizes gene assignments to functional categories while preserving the underlying data structure. The specific choice of null model depends on the biological question and the nature of potential confounding factors that must be controlled for in the analysis.
Application in Genomic Signal Detection
In the context of leadered and leaderless gene research, shuffling tests have been instrumental in validating the statistical significance of identified sequence motifs and regulatory signals. For example, when identifying TA-like promoter signals associated with leaderless genes in bacterial genomes, researchers employed shuffling tests to distinguish biologically meaningful signals from random occurrences [2]. The algorithm classified genes into SD-led, TA-led, and atypical categories based on the most probable signal in their upstream sequences, with TA-like signals approximately 10 bp upstream of the translation initiation site indicating leaderless genes.
The validation process involved comparing the observed number of TA-led genes against a null distribution generated by applying the same detection algorithm to sequences that had been shuffled while preserving dinucleotide frequencies. This approach confirmed that the identified TA-led genes significantly exceeded chance expectations, with the shuffling test demonstrating that fewer than 400 TA-led genes would be identified in randomized sequences compared to the 1,469 actually detected in the Streptomyces coelicolor A3(2) genome (p < 0.05) [2]. This rigorous statistical validation provided confidence that the detected signals represented biologically meaningful patterns rather than algorithmic artifacts or random noise.
Experimental Protocols for Shuffling Tests
Protocol 1: Sequence-Based Shuffling for Motif Validation
Purpose: To validate whether identified sequence motifs upstream of leaderless genes occur more frequently than expected by chance.
Materials:
- Genomic sequences of interest (typically 20-50 bp upstream of translation start sites)
- Motif detection algorithm (e.g., based on position weight matrices)
- Custom scripting environment (e.g., Python, R) for implementing shuffling algorithm
Methodology:
- Extract upstream sequences for all genes in the genome of interest, using annotated translation start sites as reference points.
- Apply signal detection algorithm to identify putative regulatory motifs (e.g., TA-like signals for leaderless genes).
- Record the observed count of genes containing statistically significant matches to the motif.
- Generate null distribution by creating multiple shuffled versions of the upstream sequences while preserving:
- Dinucleotide frequencies (to maintain local sequence composition)
- Sequence length distribution
- Apply the same signal detection algorithm to each shuffled sequence set and record the count of significant matches for each iteration.
- Calculate empirical p-value as (S + 1)/(N + 1), where S is the number of shuffled datasets with counts exceeding the observed count, and N is the total number of shuffling iterations.
- Apply multiple testing correction if evaluating multiple motifs simultaneously.
Interpretation: A significant p-value (typically < 0.05) indicates that the observed motif enrichment is unlikely to occur by chance alone, supporting its biological relevance.
Protocol 2: Label Shuffling for Functional Enrichment
Purpose: To test whether leaderless genes are significantly enriched for specific functional categories or experimental conditions.
Materials:
- Gene set annotations (e.g., functional categories, expression patterns)
- Classification of genes as leadered or leaderless
- Statistical computing environment
Methodology:
- Calculate observed enrichment statistic (e.g., odds ratio, hypergeometric test score) for the association between leaderless classification and functional category of interest.
- Randomly shuffle the "leaderless" label across genes while preserving:
- The total number of leaderless genes
- Potential confounding factors (e.g., genomic location, expression level)
- Recalculate the enrichment statistic for each shuffled dataset.
- Repeat the shuffling process a sufficient number of times (typically 1,000-10,000 iterations) to construct a stable null distribution.
- Determine statistical significance by comparing the observed statistic to the null distribution.
- Account for multiple comparisons when testing multiple functional categories.
Interpretation: Significant results indicate non-random association between gene leadership status and functional classification, suggesting potential biological specialization of different translation initiation mechanisms.
Table: Shuffling Test Types and Their Applications in Gene Research
| Shuffling Type | Application Context | Preservation Constraints | Null Hypothesis |
|---|---|---|---|
| Sequence Shuffling | Motif discovery, signal detection | Nucleotide composition, sequence length | Signals occur at random genomic positions |
| Label Shuffling | Functional enrichment, association studies | Number of genes in each category, gene properties | No association between leadership status and function |
| Network Shuffling | Protein-protein interaction networks | Degree distribution, network topology | Connectivity patterns occur randomly |
Visualization of Methodological Workflows
Statistical Validation Workflow for Genomic Signals
Title: Statistical validation workflow for genomic signal detection using shuffling tests
Experimental Design for Leadered/Leaderless Gene Analysis
Title: Experimental design for comparative analysis of leadered and leaderless genes
Table: Essential Research Reagents and Computational Tools for Shuffling Tests
| Resource Category | Specific Examples | Function/Purpose |
|---|---|---|
| Genomic Data Sources | NCBI GenBank, Ensembl Bacteria | Provide annotated genome sequences for signal detection |
| Sequence Analysis Tools | MEME Suite, HMMER | Identify conserved motifs in upstream regions |
| Statistical Computing | R/Bioconductor, Python SciPy | Implement shuffling algorithms and calculate significance |
| Specialized Algorithms | Di-nucleotide preserving shufflers, Position-specific scoring matrices | Generate appropriate null models for specific research questions |
| Visualization Platforms | ggplot2, Matplotlib, Graphviz | Create publication-quality figures and workflow diagrams |
| Validation Datasets | Experimentally confirmed leaderless genes (e.g., from S. coelicolor) | Benchmark computational predictions against biological truth |
Interpretation Guidelines and Reporting Standards
Critical Analysis of Shuffling Test Results
Proper interpretation of shuffling test outcomes requires careful consideration of both statistical and biological factors. A statistically significant result (typically p < 0.05) indicates that an observed pattern is unlikely to occur by random chance alone, but does not automatically imply biological importance or mechanistic relevance. Researchers should consider the effect size alongside statistical significance, as large genomic datasets may produce statistically significant results with minimal biological impact due to high statistical power.
When interpreting shuffling tests in leadered/leaderless gene research, contextual factors including genomic GC content, operon organization, and phylogenetic relationships should be considered. For example, the proportion of leaderless genes shows substantial variation across bacterial taxa, with higher representation in Actinobacteria and Deinococcus-Thermus compared to other phyla [2]. This phylogenetic signal should be accounted for when making cross-species comparisons or evolutionary inferences.
Common Pitfalls and Mitigation Strategies
Several methodological pitfalls can compromise the validity of shuffling tests in genomic research:
Inappropriate null models: Shuffling procedures that fail to preserve key sequence properties (e.g., dinucleotide frequency) may produce unrealistic null distributions, leading to inflated significance estimates. Solution: Implement conservative shuffling algorithms that preserve relevant sequence characteristics.
Multiple testing burden: Genome-scale analyses typically involve testing thousands of hypotheses simultaneously, dramatically increasing false discovery rates. Solution: Apply rigorous multiple testing corrections (e.g., Bonferroni, Benjamini-Hochberg) and report both corrected and uncorrected p-values.
Confounding factors: Unaccounted variables such as gene length, expression level, or genomic location may create spurious associations. Solution: Implement stratified shuffling approaches or include covariates in the analytical model.
Computational intensity: Comprehensive shuffling tests with large genomic datasets may require substantial computational resources. Solution: Utilize efficient algorithms, parallel computing, and appropriate subsampling strategies when necessary.
Shuffling tests provide an essential statistical framework for validating genomic signals in leadered and leaderless gene research, enabling robust distinction between biologically meaningful patterns and random noise. As genomic datasets continue to expand in both size and complexity, these non-parametric approaches will play an increasingly important role in ensuring the reliability of biological inferences.
Future methodological developments will likely focus on enhancing the sophistication of null models to better account for genomic architecture, improving computational efficiency for large-scale applications, and integrating shuffling tests with other statistical approaches to provide comprehensive validation frameworks. Additionally, as single-cell sequencing and other emerging technologies reveal new dimensions of transcriptional complexity, shuffling tests will need to adapt to address novel analytical challenges in characterizing translation initiation mechanisms.
The integration of rigorous statistical validation through shuffling tests with experimental molecular biology approaches will continue to drive advances in our understanding of leadered and leaderless genes, ultimately illuminating the evolutionary dynamics and functional implications of alternative translation initiation mechanisms across the bacterial domain.
The study of translation initiation mechanisms in prokaryotes reveals a fundamental dichotomy between leadered and leaderless genes. Leadered genes possess a 5'-untranslated region (5'-UTR) that typically contains a Shine-Dalgarno (SD) sequence, which guides the ribosome to the initiation site through complementary base pairing with the 16S rRNA [2]. In contrast, leaderless genes completely lack a 5'-UTR, with the start codon positioned at or very near the transcription start site [2]. This structural distinction implies different mechanistic strategies for ribosome recruitment and translation initiation, making accurate experimental validation crucial for understanding gene regulation.
While bioinformatic analyses have revealed that leaderless genes are "widespread, although not dominant, in a variety of bacteria" and can constitute over twenty percent of genes in certain phyla like Actinobacteria [2], computational predictions alone cannot confirm functional translation initiation mechanisms. Experimental verification through fluorescence reporter systems and high-resolution transcription start site (TSS) mapping provides the necessary empirical evidence to distinguish between these initiation strategies, validate bioinformatic predictions, and understand the regulatory implications of each mechanism in different biological contexts.
Core Principles: Leadered versus Leaderless Genes
Molecular Signatures and Identification Criteria
The structural and sequence differences between leadered and leaderless genes create distinct molecular signatures that guide both computational prediction and experimental design.
Table 1: Key Characteristics of Leadered versus Leaderless Genes
| Feature | Leadered Genes | Leaderless Genes |
|---|---|---|
| 5'-UTR | Present (typically 20-50+ nucleotides) | Absent or extremely short |
| SD Sequence | Present upstream of start codon | Absent |
| Start Codon Context | AUG, GUG, UUG; preceded by SD | AUG, GUG, UUG; at or near TSS |
| Promoter Position | Upstream of TSS, which precedes coding sequence | Upstream of start codon/TSS |
| Initiation Mechanism | SD-mediated ribosome binding | Direct ribosome binding to start codon |
| Evolutionary Distribution | Widespread across bacteria | Varies significantly (≥20% in Actinobacteria) [2] |
Computational Identification and Bioinformatic Evidence
Bioinformatic approaches for identifying leaderless genes typically involve analyzing sequences upstream of annotated start codons. Research across 953 bacterial genomes has demonstrated that TA-like signals located approximately 10-12 bp upstream of the translation initiation site (corresponding to the -10 promoter element) serve as reliable indicators of leaderless architecture in bacteria [2]. These signals exhibit a consensus pattern of TANNNT, resembling the -10 box of σ70 factor binding sites [2].
Statistical validation is essential for distinguishing genuine signals from random occurrence. Shuffling tests that preserve dinucleotide frequency have demonstrated that the number of TA-led genes identified in bacterial genomes significantly exceeds what would be expected by chance, providing statistical confidence in these classifications [2]. For example, in Streptomyces coelicolor A3(2), 1,469 genes (18.9%) were identified as leaderless through this computational approach, with only 400 expected by random chance [2].
Fluorescence Reporter Systems for Validation
Reporter Design Principles and Considerations
Fluorescence reporter systems enable quantitative assessment of gene expression by linking regulatory sequences to easily measurable fluorescent proteins. These systems function as transcriptional biosensors that report on the activity of upstream regulatory elements in living cells and organisms [40].
The core design involves placing the putative regulatory sequence (promoter and 5'-UTR, if present) upstream of a fluorescent protein coding sequence. For leaderless gene validation, the critical design element is ensuring that the start codon of the fluorescent protein constitutes the first translated codon, immediately following any regulatory elements. This architecture mirrors the native structure of leaderless transcripts and prevents the introduction of artificial 5'-UTRs that would convert a leaderless context into a leadered one.
Advanced Reporter Biosensor: CLEARoptimized Case Study
Recent methodological advances have produced sophisticated reporter systems like the CLEARoptimized biosensor, designed to study transcription factors but exemplifying optimal reporter design principles applicable to leaderless/leadered gene validation [40].
Table 2: Components of the CLEARoptimized Fluorescence Reporter System
| Component | Description | Function in Validation |
|---|---|---|
| Synthetic Promoter | 6 coordinated lysosomal expression and regulation (CLEAR) motifs [40] | Contains multiple TFEB/TFE3 binding sites; can be adapted for bacterial promoters |
| Minimal Promoter | Thymidine kinase (Tk) promoter | Provides basal transcription machinery interaction |
| Reporter Genes | Luciferase (luc2) and tdTomato fluorescent protein separated by T2A peptide [40] | Dual reporting enables normalization and quantification in different modalities |
| T2A Self-proteolytic Peptide | 22-amino acid peptide sequence [40] | Enables coordinated expression of both reporters from single transcript |
This biosensor was specifically engineered through in-depth bioinformatic analysis of 128 TFEB-target genes, which revealed that optimal responsive elements are "typically clustered in multiple copies, more frequently located within -200 base pairs from the transcription start site" [40]. The synthetic promoter was computationally validated using JASPAR2020 to reduce off-target transcription factor binding, resulting in a highly specific reporter system [40].
Experimental Protocol: Fluorescence Reporter Validation
Step 1: Vector Construction
- Clone the putative regulatory sequence (approximately 200-300 bp upstream of the start codon, or full 5' region for leaderless genes) into a fluorescence reporter vector.
- For leaderless gene validation, ensure precise positioning so the start codon of the fluorescent reporter is the first codon following the promoter.
- Use restriction enzyme cloning or Gibson assembly to insert the fragment between the promoter and reporter gene, eliminating any endogenous 5'-UTR from the vector backbone.
Step 2: Cell Transformation and Culture
- Transform the constructed plasmid into an appropriate bacterial strain (e.g., E. coli for general studies or specific species like Streptomyces for native context).
- Culture transformed cells under standardized conditions, with appropriate controls (empty vector, positive control with known leaderless promoter, etc.).
- For in vivo imaging in bacteria, use specialized equipment capable of detecting bacterial fluorescence above autofluorescence.
Step 3: Signal Measurement and Quantification
- Measure fluorescence intensity using a microplate reader or fluorescence microscope with appropriate filter sets for the fluorescent protein (e.g., 554/581 nm excitation/emission for tdTomato).
- Normalize fluorescence values to cell density (OD600) to account for growth differences.
- For dual-reporter systems like CLEARoptimized, use luciferase activity as a primary quantitative measure and fluorescence for localization and single-cell analysis [40].
Step 4: Data Interpretation
- Compare expression levels from test constructs to appropriate controls.
- Leaderless constructs typically show constitutive but potentially lower expression compared to optimized leadered genes.
- Significant fluorescence above background confirms functional translation initiation, while absence suggests the sequence may not function as a true leaderless promoter in the experimental context.
Transcription Start Site (TSS) Mapping Methodologies
Principles of TSS Determination
TSS mapping provides direct experimental evidence for classifying genes as leadered or leaderless by precisely identifying the 5' end of transcripts. If the TSS corresponds exactly to the first nucleotide of the start codon, the gene is definitively leaderless. A TSS located upstream of the start codon creates a 5'-UTR and indicates a leadered architecture.
High-resolution TSS mapping typically employs specialized RNA sequencing techniques that capture the 5' ends of transcripts, providing genome-wide data that can classify initiation mechanisms for all genes in a prokaryotic genome.
High-Throughput TSS Mapping Protocol
Step 1: RNA Harvesting and Quality Control
- Grow bacterial cultures to mid-log phase under experimental conditions of interest.
- Harvest cells and extract total RNA using a hot phenol protocol or commercial kit optimized for bacterial RNA.
- Treat with DNase I to remove genomic DNA contamination.
- Assess RNA quality using bioanalyzer or agarose gel electrophoresis; ensure RIN (RNA Integrity Number) >8.0 for optimal results.
Step 2: 5' RNA Adapter Ligation
- Dephosphorylate RNA using calf intestinal phosphatase to remove 5' phosphates from degraded fragments.
- Treat with tobacco acid pyrophosphatase (TAP) to remove 5' caps from eukaryotic RNA (less critical for bacterial work but included for completeness).
- Ligate a specialized RNA adapter to the 5' ends of transcripts using T4 RNA ligase.
- This adapter contains a priming site for reverse transcription and serves as a universal 5' anchor in subsequent amplification steps.
Step 3: cDNA Library Preparation and Sequencing
- Reverse transcribe the adapter-ligated RNA using random hexamers or gene-specific primers.
- Amplify the cDNA library using PCR with barcoded primers to enable multiplexing.
- The amplicons contain the barcode, adapter sequence, and the 5' end of the original transcript, enabling precise mapping of TSS positions.
- Sequence the library using Illumina or similar high-throughput platforms, aiming for 3-10 million reads per sample depending on genome complexity.
Step 4: Bioinformatic Analysis and TSS Classification
- Process raw sequencing data to trim adapter sequences and quality filter reads.
- Align processed reads to the reference genome using specialized tools like FASTQINS for transposon insertion sites or standard aligners like Bowtie2 for RNA-seq data [41].
- Identify TSS positions as genomic coordinates with significant enrichment of 5' read starts.
- Classify genes based on the distance between TSS and start codon:
- Leaderless: TSS corresponds to start codon position (distance = 0)
- Leadered: TSS located upstream of start codon (distance > 0)
- Uncertain: Multiple TSSs or ambiguous mapping
Integrated Data Analysis and Interpretation
Correlating Fluorescence and TSS Data
The combination of fluorescence reporter assays and TSS mapping creates a powerful validation framework where results from both methods should converge to support the same classification.
Table 3: Expected Experimental Outcomes for Leadered vs. Leaderless Genes
| Method | Leaderless Gene Pattern | Leadered Gene Pattern |
|---|---|---|
| Fluorescence Reporter | Moderate fluorescence, less dependent on SD mutations | Strong fluorescence, dependent on intact SD sequence |
| TSS Mapping | TSS corresponds precisely to start codon position | TSS located upstream of start codon (creating 5'-UTR) |
| Sequence Analysis | TA-like signal ~10 bp upstream; no SD motif | SD sequence 5-10 bp upstream of start codon |
| Evolutionary Context | More common in certain bacterial phyla (e.g., Actinobacteria) [2] | Dominant mechanism in most bacterial species |
When discrepancies occur between fluorescence and TSS data (e.g., TSS suggests leaderless architecture but fluorescence is absent), consider alternative explanations:
- The gene may require specific regulatory factors absent in the experimental system
- mRNA secondary structure might block translation despite correct TSS
- The start codon annotation might be incorrect
- Post-transcriptional regulation might influence outcomes
Statistical Validation and Threshold Determination
Robust statistical analysis is essential for confident classification. For TSS mapping, establish minimum read count thresholds (typically >1000 mapped reads at a position) to distinguish true TSS from background noise [42]. For fluorescence data, apply appropriate statistical tests (t-tests, ANOVA) to ensure significant differences between test constructs and controls.
Shuffling tests that preserve dinucleotide composition provide a valuable negative control for computational predictions. In one comprehensive study, this approach demonstrated that the number of identified TA-led genes in bacterial genomes (indicating leaderless architecture) significantly exceeded random expectation, with observed counts typically 3-4 times higher than background [2].
The Scientist's Toolkit: Essential Research Reagents
Table 4: Key Research Reagent Solutions for Validation Experiments
| Reagent/Category | Specific Examples | Function/Application |
|---|---|---|
| Reporter Vectors | pCLEARoptimized, pGL4-series, custom T7-based vectors | Backbone for constructing fluorescence/luminescence reporter fusions |
| Fluorescent Proteins | tdTomato, GFP, mCherry, YFP | Visual reporters for gene expression and localization studies |
| Enzymes for Library Prep | T4 RNA Ligase, Tobacco Acid Pyrophosphatase, Polynucleotide Kinase | Essential for 5' adapter ligation in TSS mapping protocols |
| High-Throughput Sequencers | Illumina NovaSeq, MiSeq, PacBio Revio | Platform for TSS mapping and transcriptome analysis |
| Cell Lines/Strains | HEK293, HeLa, E. coli BW25113, B. subtilis 168 | Model systems for heterologous and homologous expression |
| Bioinformatic Tools | JASPAR2020, FASTQINS, Bowtie2, custom Perl/Python scripts | Computational analysis of promoter motifs and NGS data |
The integrated application of fluorescence reporter systems and TSS mapping provides a robust experimental framework for validating translation initiation mechanisms and distinguishing between leadered and leaderless genes. These methodologies transform computational predictions into biologically verified mechanisms, enabling researchers to move beyond sequence analysis to functional characterization.
As research in this field advances, the development of more sensitive fluorescent proteins, improved high-throughput sequencing methods, and sophisticated computational models will further enhance our ability to precisely categorize and understand the functional significance of different translation initiation strategies across diverse biological systems. The experimental approaches detailed in this technical guide provide a foundation for these future investigations, establishing standardized methodologies that will facilitate comparative analyses and deepen our understanding of gene regulation evolution in prokaryotic systems.
The central dogma of molecular biology outlines the flow of genetic information from DNA to RNA to protein. However, the relationship between these layers is not always linear or straightforward. Global profiling through integrated transcriptomics and proteomics has emerged as a powerful approach to comprehensively map these complex relationships and experimentally validate gene models. This integrated approach, often termed proteogenomics, is particularly crucial for refining genome annotations and understanding the nuanced expression dynamics between different gene classes, most notably leadered and leaderless genes [43] [44].
The distinction between these gene structures is a fundamental aspect of transcriptional and translational regulation. Leadered genes conform to the canonical initiation model, where a 5' untranslated region (5' UTR) contains regulatory elements, typically a Shine-Dalgarno (SD) sequence, that guides ribosome binding. In contrast, leaderless genes lack a 5' UTR entirely, with the transcription start site residing at or very near the start codon, thus employing non-canonical, and in some cases ancient, initiation mechanisms [10] [17] [14]. While once considered a rarity in bacteria, large-scale genomic and transcriptomic studies have revealed that leaderless genes are widespread, comprising a significant portion of the transcriptome in certain taxa, including approximately 14% of genes in Mycobacterium tuberculosis and over 20% in Actinobacteria and Deinococcus-Thermus [10] [13] [14]. This technical guide provides a detailed framework for using integrated proteomics and transcriptomics to analyze gene expression, with a specific focus on the technical considerations for studying these distinct gene structures.
Fundamental Differences Between Leadered and Leaderless Genes
Understanding the core structural and mechanistic differences between leadered and leaderless genes is a prerequisite for designing effective profiling experiments. The table below summarizes the key characteristics that influence their expression and regulation.
Table 1: Key Characteristics of Leadered and Leaderless Genes
| Feature | Leadered Genes | Leaderless Genes |
|---|---|---|
| 5' UTR | Present, often 40-60 nucleotides long [13] | Absent or very short [10] [14] |
| Shine-Dalgarno (SD) Sequence | Typically present upstream of start codon [10] | Absent [14] |
| Primary Translation Initiation Mechanism | 30S ribosomal subunit binding via SD-anti-SD interaction; factor-assisted [14] | Direct binding by 70S ribosomes; can be factor-independent or use IF2/eIF5B [17] [14] |
| Conservation | Considered the dominant, canonical mechanism in prokaryotes [10] | Abundant in Archaea and specific bacterial phyla; hypothesized ancient mechanism [10] [17] |
| Impact on mRNA Stability | 5' UTR secondary structure can influence mRNA half-life [13] | Often shorter mRNA half-life, potentially due to lack of 5' UTR protection [13] |
| Response to Cellular Stress | Often downregulated during stress (e.g., eIF2α phosphorylation) [17] | Can be relatively resistant to specific stresses that inhibit canonical initiation [17] |
These fundamental differences mean that the two gene classes often behave differently in expression analyses. For instance, the lack of a 5' UTR in leaderless transcripts means that standard gene-finding algorithms, which often rely on detecting SD sequences, may misannotate or entirely miss them [10] [14]. Furthermore, the coupling of transcription and translation, a key feature in prokaryotes, is disrupted for leaderless genes, which can impact mRNA turnover rates [13]. Consequently, integrated multi-omics approaches are not merely beneficial but essential for accurate annotation and expression analysis.
Experimental Design and Workflow for Integrated Profiling
A robust proteogenomic workflow involves the coordinated application of high-throughput transcriptomic and proteomic techniques to refine genome annotation and quantify expression. The following diagram outlines a generalized, high-level workflow for this process.
Diagram 1: High-level Proteogenomic Workflow for Gene Model Refinement
Transcriptome Profiling for Gene Structure Elucidation
The first pillar of the integrated approach involves deep transcriptome sequencing to map the precise boundaries of transcripts.
- RNA Sequencing (RNA-seq): Deep sequencing of total RNA provides a comprehensive view of the transcriptome. This data is crucial for identifying expressed regions of the genome, confirming exon-intron structures in eukaryotes, and detecting novel transcripts that may have been missed by computational prediction [43]. For prokaryotes, it confirms the expression of putative genes.
- Translation Start Site (TSS) Mapping: Specifically defining the 5' end of transcripts is critical for distinguishing leadered from leaderless genes. Techniques like RACE (Rapid Amplification of cDNA Ends) or specialized RNA-seq protocols for enriched 5' ends enable precise identification of the transcription start site. A TSS coinciding with the start codon (ATG, GTG, etc.) is a definitive marker of a leaderless gene [14].
- Ribosome Profiling (Ribo-seq): This technique involves the deep sequencing of mRNA fragments protected by ribosomes. It provides a nucleotide-resolution snapshot of translation by revealing the positions of actively translating ribosomes across the transcriptome. Ribo-seq is particularly powerful for identifying novel, small open reading frames (ORFs) and for validating the translation of leaderless transcripts, as it shows ribosome occupancy directly at the 5' end of an mRNA without an upstream SD sequence [14].
Proteome Profiling for Experimental Validation
The second pillar uses mass spectrometry (MS)-based proteomics to provide direct experimental evidence for protein synthesis, which is the ultimate validation of a coding gene.
- Sample Preparation and Fractionation: To achieve broad proteome coverage, proteins are extracted from multiple biological conditions, tissues, or cell fractions. These extracts are digested into peptides, which are then fractionated using chromatography (e.g., strong cation exchange) or high-pH reversed-phase separation. This reduces sample complexity and increases the depth of analysis [43].
- Liquid Chromatography-Tandem Mass Spectrometry (LC-MS/MS): Fractionated peptides are separated by liquid chromatography and injected into a high-resolution mass spectrometer. The instrument isolates peptide ions, fragments them, and measures the mass-to-charge ratio of both the intact peptides and their fragments, generating MS/MS spectra [43] [44].
- Database Search and Novel Peptide Discovery: The acquired MS/MS spectra are searched against a protein database derived from the genomic sequence. The standard search confirms annotated proteins. The proteogenomic approach involves additional searches against customized databases that include novel coding sequences predicted from the transcriptomic data (e.g., from TSS mapping and Ribo-seq). Peptides that map to these novel regions provide robust evidence for new gene models or corrections to existing ones [43] [44].
Table 2: Key Research Reagents and Platforms for Integrated Profiling
| Category / Reagent | Specific Examples / Techniques | Primary Function in Analysis |
|---|---|---|
| Transcriptomics | RNA-seq, TSS Mapping (RACE), Ribo-seq | Maps transcript boundaries, identifies 5' ends, and profiles translational activity to define gene structures. |
| Proteomics | High-resolution Mass Spectrometer (Orbitrap), Liquid Chromatography (LC) | Separates and fragments peptides to generate identification data (MS/MS spectra). |
| Bioinformatics Tools | TopHat2 (read alignment), EuGenoSuite/ProteoAnnotator (proteogenomic search), Custom Perl/Python scripts | Aligns sequencing data, integrates disparate data types, and performs customized database searches. |
| Custom Databases | Six-frame translation, Novel ORFs from RNA-seq, N-terminal peptide database | Expands search space beyond annotated genes to discover novel proteins and correct gene models. |
| Validation Reagents | Synthetic Peptides, Reporter Gene Constructs (e.g., YFP/Luciferase) | Provides orthogonal validation for novel peptide identifications and tests cis-regulatory requirements. |
A Practical Case Study: Profiling the Mycobacterial Translational Landscape
The application of this integrated approach is powerfully illustrated by research on mycobacteria, which exhibit a high proportion of leaderless genes. A landmark study combined TSS mapping, RNA-seq, Ribo-seq, and N-terminal peptide MS to re-annotate the genomes of Mycobacterium smegmatis and Mycobacterium tuberculosis [14].
The experimental protocol can be summarized as follows:
- Define Transcript Structures: TSS mapping was performed genome-wide to identify thousands of transcription start sites. This revealed that approximately 24% of mycobacterial transcripts are leaderless, starting with the first nucleotide of the start codon [14].
- Profile Translational Activity: Ribo-seq was employed to map the positions of ribosomes across the M. smegmatis transcriptome. This data showed robust ribosome occupancy at the 5' end of leaderless transcripts, confirming their active translation. It also identified hundreds of small, unannotated ORFs at the 5' ends of transcripts, suggesting a previously hidden layer of the proteome [14].
- Validate Protein Expression: N-terminal proteomics using MS provided direct experimental evidence for the translation of these leaderless ORFs. peptides matching the precise N-termini of proteins, including many that were newly discovered, confirmed their existence and allowed researchers to distinguish the true start codon from internal sites [14].
- Functional Validation with Reporters: The study used a fluorescent reporter system (YFP) to mechanistically probe the requirements for translation initiation. By systematically mutating the start codon and 5' sequence, they demonstrated that an ATG or GTG at the 5' end is both necessary and sufficient for robust leaderless translation in mycobacteria [14].
This integrated workflow led to the discovery of hundreds of new small proteins and fundamentally refined the understanding of the mycobacterial translational landscape, demonstrating that leaderless initiation is a major, robust mechanism in these organisms.
Critical Data Analysis and Visualization Strategies
The volume of data generated from transcriptomic and proteomic pipelines requires careful analysis and clear visualization to draw meaningful conclusions, particularly when comparing different gene classes.
Quantitative Comparison of Gene Classes
Integrated profiling allows for the systematic comparison of expression metrics between leadered and leaderless genes. The following table summarizes potential findings from such an analysis.
Table 3: Hypothetical Comparative Expression Data from an Integrated Profiling Study
| Expression Metric | Leadered Genes | Leaderless Genes | Technical Implication |
|---|---|---|---|
| Protein-to-mRNA Ratio (Median) | ~1.0 (Reference) | ~0.8 - 1.2 [13] | No systematic, major difference in average translation efficiency. |
| Median mRNA Half-life | Longer (e.g., >5 min) [13] | Shorter (e.g., <5 min) [13] | Leaderless transcripts may be less stable; requires careful normalization in RNA-seq. |
| Predicted Transcript Production Rate | Variable | Can be higher for some [13] | Suggests compensation for shorter half-life to maintain protein output. |
| Proteogenomic Validation Rate | High for annotated genes | Lower for previously annotated genes; high for novel 5' ORFs [14] | Standard annotation pipelines are biased against leaderless genes. |
Visualizing Experimental Outcomes and Workflows
Effective diagrams are essential for communicating the logic of experimental designs and the biological insights gained. The diagram below illustrates the core finding from the mycobacterial case study, showing how multi-omics data converges to define a leaderless gene.
Diagram 2: Multi-omics Validation of a Leaderless Gene
Integrated proteogenomic analysis provides an unparalleled, empirical framework for understanding gene expression. By moving beyond in silico predictions and combining the power of transcriptomics with the validating force of proteomics, researchers can achieve a more accurate and comprehensive picture of the genomic landscape. This approach is indispensable for studying non-canonical gene structures like leaderless genes, which are often misannotated yet play critical biological roles. As these methodologies become more accessible and standardized, their application will continue to refine genome annotations across the tree of life, drive discoveries of novel genetic elements, and deepen our understanding of the fundamental mechanisms governing gene expression in all its forms.
Navigating Experimental Complexities in Leaderless Gene Research
Accurate annotation of the Translation Initiation Site (TIS) represents a fundamental challenge in genomic studies, with profound implications for predicting protein sequences, understanding gene regulation, and facilitating drug discovery efforts. The selection of the correct start codon is complicated by biological realities that computational methods must address: multiple potential initiation codons exist within a single genomic context, and the mechanisms governing initiation differ significantly between leadered genes (with 5' untranslated regions) and leaderless genes (lacking 5' UTRs) [2]. This technical guide examines the strategies developed to resolve start codon ambiguity, framed within the critical research distinction between leadered and leaderless gene structures.
The significance of precise TIS annotation extends far beyond correct protein prediction. Inaccurate initiation site identification can misdirect operon predictions, promoter identification, and the discovery of small non-coding RNAs, as these analyses frequently depend on accurate intergenic distance calculations [45]. Within the context of drug development, understanding these fundamental genetic regulatory mechanisms provides crucial insights for targeting pathogenic bacteria, particularly those like Mycobacterium species that utilize both initiation mechanisms [27].
Biological Foundations: Leadered vs. Leaderless Genes
Molecular Mechanisms of Translation Initiation
Prokaryotic translation initiation occurs through two principal pathways, each with distinct sequence requirements and molecular machinery.
Leadered Gene Initiation: Leadered genes contain 5' untranslated regions (5'-UTRs) that harbor a ribosome binding site (RBS), most commonly the Shine-Dalgarno (SD) sequence with consensus 5'-AGGAGG-3' [46]. This sequence base-pairs with the anti-Shine-Dalgarno sequence at the 3' end of the 16S rRNA component of the 30S ribosomal subunit, facilitating proper positioning of the ribosome at the initiation codon [46]. The optimal spacing between the SD sequence and the start codon is typically 3-8 nucleotides, though this distance exhibits species-specific variation [45].
Leaderless Gene Initiation: Leaderless genes completely lack 5'-UTRs, with the transcription start site coinciding directly with the first nucleotide of the initiation codon [2]. Without an upstream RBS to facilitate ribosomal binding, initiation depends primarily on the start codon itself and potentially downstream sequence elements, employing a mechanism that bears similarity to eukaryotic initiation [2].
Table 1: Comparative Features of Leadered and Leaderless Genes
| Feature | Leadered Genes | Leaderless Genes |
|---|---|---|
| 5' UTR | Present (variable length) | Absent |
| RBS/SD Sequence | Present (highly conserved to degenerate) | Absent |
| Initiation Mechanism | SD-antiSD base pairing | Start codon recognition, potentially eukaryotic-like |
| Start Codon Preference | Varies by species (ATG, GTG, TTG) | Primarily ATG |
| Prevalence in Bacteria | Variable across taxa | Widespread but not dominant (up to >20% in Actinobacteria) |
| Evolutionary Trend | Increasing prevalence | Decreasing prevalence in bacterial evolution |
While ATG represents the most efficient and commonly used start codon across prokaryotes, significant species-specific variation exists in the utilization of alternative initiation codons. Research comparing Escherichia coli and Bacillus subtilis reveals distinct preferences: in E. coli, GTG represents the predominant alternative start codon (7.2%), while B. subtilis shows stronger preference for TTG (10.7%) over GTG (8.6%) [45]. This variation in codon preference presents a considerable challenge for computational TIS prediction algorithms trained on model organisms like E. coli when applied to diverse bacterial species.
Computational Prediction Methodologies
Integrated Bayesian Approaches
The Hon-yaku methodology represents a biology-driven Bayesian approach that integrates multiple sequence features into a unified scoring function for TIS prediction [45]. This supervised learning framework combines six biologically-relevant elements to achieve prediction accuracies exceeding 90% in both E. coli and B. subtilis datasets:
- RBS sequence motif and its positional relationship to potential start codons
- Distance between putative TIS and RBS sequence (typically 3-8 nucleotides)
- Nucleotide composition of the start codon (accounting for species-specific preferences)
- A-rich sequences immediately downstream of the start codon, which may prevent inhibitory RNA secondary structures
- Expected protein length distribution based on genomic context
- Operon orientation and its effect on intergenic distance distributions [45]
This integrated approach demonstrates particularly improved performance in GC-rich organisms like Pseudomonas aeruginosa and Burkholderia pseudomallei, where traditional methods struggle with false positives due to statistically generated long ORFs [45].
Signal-Based Classification Algorithms
Advanced computational methods have been developed to classify genes based on initiation signals through multi-signal analysis of upstream regions. One such algorithm examines 20-basepair TIS upstream sequences in bacteria to categorize genes into three classes:
- SD-led genes: Contain identifiable Shine-Dalgarno sequences
- TA-led genes: Feature TA-like signals approximately 10-12 bp upstream of TIS, indicating leaderless transcription with promoter elements resembling the σ70 factor -10 box "TATAAT" consensus
- Atypical genes: Lack clear SD or TA signals, potentially employing alternative initiation mechanisms [2]
This classification system has revealed the widespread distribution of leaderless genes across diverse bacterial taxa, with particularly high prevalence in Actinobacteria and Deinococcus-Therpus, where over twenty percent of genes utilize leaderless initiation [2].
Table 2: Performance Comparison of TIS Prediction Methods
| Method | Approach | E. coli Accuracy | B. subtilis Accuracy | GC-rich Organisms |
|---|---|---|---|---|
| Hon-yaku | Bayesian integrated features | 93.2% | 92.7% | Improved performance |
| Glimmer | Markov models, longest ORF | Lower 5'-end accuracy | Reduced accuracy | Struggles with long ORFs |
| MED-Start | Unspecified | ~90% | N/R | ~5% accuracy in high GC |
| GeneMark | Markov chains, atypical states | Moderate | Moderate | Handles horizontal transfer |
Experimental Validation Protocols
Reporter Construct Systems for Initiation Efficiency
Determining the functional impact of leadered versus leaderless structures requires carefully controlled experimental systems. One robust methodology employs fluorescence reporter constructs to measure multiple expression parameters simultaneously:
Protocol: Multiparameter Reporter Assay
Construct Design: Clone candidate 5' UTR sequences (for leadered genes) or full leaderless initiation regions upstream of a fluorescent protein reporter gene (e.g., GFP) [27]
Control Elements: Include standardized synthetic 5' UTR controls to establish baseline expression profiles
Expression Measurement:
- Protein Abundance: Quantify fluorescence intensity as a proxy for translation efficiency
- mRNA Abundance: Extract RNA and perform quantitative RT-PCR to determine transcript levels
- mRNA Half-life: Treat cultures with transcription inhibitors and track transcript disappearance over time to calculate stability [27]
Transcript Production Rate Calculation: Derive relative transcription rates by combining mRNA abundance and stability data
This integrated approach revealed that the sigA 5' UTR in Mycobacterium smegmatis confers increased transcript production rate but shorter mRNA half-life compared to synthetic controls, while leaderless transcripts exhibited similar translation efficiency but lower predicted production rates [27].
Genome-Wide Computational Validation
For large-scale validation of TIS predictions, statistical significance testing provides essential confirmation of identified initiation signals:
Protocol: Shuffling Test for Signal Significance
Generate Null Sequences: Create randomized versions of native upstream sequences while preserving dinucleotide frequency to maintain local sequence composition biases [2]
Signal Detection: Apply the same signal detection algorithm (e.g., for TA-like motifs) to both native and shuffled sequences
Statistical Comparison: Calculate the excess of signals in native sequences compared to shuffled controls
Confidence Thresholding: Establish minimum threshold values for significant signal identification based on statistical significance (e.g., p < 0.01) [2]
This method demonstrated that among 7,769 protein-coding genes in Streptomyces coelicolor A3(2), 1,469 (18.9%) contained statistically significant TA-like signals indicative of leaderless transcription, far exceeding the <400 false positives expected by chance [2].
Visualization of Key Mechanisms and Workflows
Translation Initiation Pathways Diagram
Figure 1. Leadered vs. Leaderless Initiation Mechanisms
TIS Prediction Workflow Diagram
Figure 2. Computational TIS Prediction Pipeline
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Research Reagents for TIS Investigation
| Reagent/Solution | Function/Application | Experimental Context |
|---|---|---|
| Fluorescence Reporter Plasmids | Quantitative measurement of translation efficiency via protein abundance | Validation of putative TIS functionality [27] |
| qRT-PCR Reagents | mRNA quantification and stability assessment | Correlation of transcript levels with protein output [27] |
| Shuffled Sequence Controls | Statistical background estimation | Computational validation of signal significance [2] |
| Species-Specific Training Sets | Reference data for supervised learning | Bayesian prediction algorithms [45] |
| 16S rRNA-Targeting Reagents | Ribosomal binding studies | Investigation of SD-antiSD interactions [46] |
| Transcriptional Inhibitors | mRNA decay rate measurements | Determination of transcript half-lives [27] |
Accurate resolution of start codon ambiguity requires integrated methodologies that account for the fundamental biological distinction between leadered and leaderless gene architectures. The most successful approaches combine multiple sequence features through Bayesian statistics or similar integrative frameworks, validated through both computational significance testing and experimental reporter assays [45] [2].
Future advancements in TIS annotation will likely emerge from several promising directions: improved algorithms that better account for the complex sequence determinants of non-SD-led initiation, expanded experimental validation across diverse bacterial taxa, and single-cell approaches to resolve cell-to-cell variation in initiation events. For drug development professionals, understanding these initiation mechanisms provides crucial insights for targeting pathogenic bacteria with atypical initiation patterns, potentially revealing novel therapeutic avenues against persistent infectious agents like Mycobacterium tuberculosis [27].
The evolutionary trajectory of translation initiation mechanisms, with leaderless genes showing a decreasing proportion throughout bacterial evolution [2], suggests that ancient initiation mechanisms may persist in modern pathogens, offering both challenges and opportunities for antimicrobial strategies focused on the fundamental processes of gene expression.
In the broader context of research on leadered versus leaderless genes, the untranslated regions (UTRs) of messenger RNA (mRNA) serve as critical regulatory hubs that fine-tune gene expression. While the coding sequence determines a protein's amino acid sequence, the 5' and 3' UTRs govern the efficiency with which that protein is synthesized, influencing multiple facets of mRNA metabolism including stability, localization, and translational efficiency. The fundamental distinction between leadered genes (containing 5' UTRs) and leaderless genes (initiating directly at the start codon) represents an evolutionary divergence in translational control mechanisms with profound implications for cellular adaptation. In pathogenic bacteria like Mycobacterium tuberculosis, approximately 14% of genes are leaderless, an unusually high prevalence that suggests specialized regulatory functions for stress adaptation during infection [13]. This technical guide examines how UTR length and composition mechanistically influence translation efficiency, providing researchers with both theoretical frameworks and practical methodologies for investigating these relationships in experimental systems.
Fundamental Mechanisms of Translation Initiation
Canonical Initiation Pathways for Leadered mRNAs
Leadered mRNAs, which constitute the majority of transcripts in most organisms, rely on structured initiation mechanisms that begin with ribosome dissociation. In bacteria, the Shine-Dalgarno sequence within the 5' UTR base-pairs with the 16S rRNA of the 30S ribosomal subunit, facilitating proper positioning of the start codon [47]. This process typically requires dissociation of the 70S ribosome into 50S and 30S subunits before initiation can proceed. In eukaryotes, a more complex initiation pathway involves the scanning 40S ribosomal subunit, multiple initiation factors (eIFs), and recognition of the 5' cap structure [17]. The secondary structure of 5' UTRs in leadered transcripts plays a critical regulatory role in these processes, as extensive structure can impede ribosomal scanning and translation initiation [48].
Specialized Initiation Mechanisms for Leaderless mRNAs
Leaderless mRNAs employ fundamentally different initiation mechanisms that bypass many conventional requirements. Research across diverse biological systems reveals that leaderless transcripts can bind directly to non-dissociated ribosomes—70S in bacteria [49] and 80S in eukaryotes [17]—without the need for canonical initiation factors. This ancient initiation pathway demonstrates remarkable flexibility, with studies in mammalian systems revealing at least four distinct initiation mechanisms available to leaderless mRNAs: 80S-mediated, eIF2-dependent, eIF2D-mediated, and eIF5B/IF2-assisted initiation [17]. The factor-independent mechanism is particularly significant as it provides translational resistance to various cellular stresses that impair standard initiation pathways.
Quantitative Analysis of UTR Impact on Translation
5' UTR Length and Composition Effects
The length of the 5' UTR significantly influences translational regulation, with both extremely short and excessively long UTRs generally impairing efficiency. In mycobacteria, the median 5' UTR length is approximately 48-56 nucleotides, but substantial variation exists [13]. Research demonstrates that specific nucleotide composition also critically impacts translation. In Escherichia coli, systematic libraries of 5' UTR sequences revealed that regions lacking cytosine (C) nucleotides showed enhanced translation efficiency, suggesting that nucleotide composition independently influences efficiency beyond length considerations [50]. The secondary structure near the 5' cap site additionally plays a recognized role in microRNA-mediated gene regulation in animals, with targeted mRNAs exhibiting increased local structure in this region [48].
3' UTR Length and Regulatory Functions
The 3' UTR length substantially influences both translational efficiency and mRNA stability, with particularly pronounced effects on poly(A)- transcripts. In mammalian cells, increasing 3' UTR length from 4 to 104 bases enhanced translational efficiency by 38-fold for poly(A)- mRNAs [51]. The stimulatory effect of poly(A) tail addition diminished from 97-fold to only 2.3-fold when 3' UTR length increased from 19 to 156 bases, indicating an interaction between poly(A) tail function and 3' UTR length [51]. Recent research on mRNA therapeutics has further demonstrated that engineered AU-rich elements in 3' UTRs can enhance both stability and translation through interactions with RNA-binding proteins like HuR [52].
Comparative Features of Leadered vs. Leaderless Genes
Table 1: Characteristics of Leadered and Leaderless Translation Systems
| Feature | Leadered Genes | Leaderless Genes |
|---|---|---|
| Prevalence | ~86% of genes in mycobacteria [13] | ~14% of genes in mycobacteria [13] |
| Initiation Mechanism | Ribosome dissociation required; SD-dependent (bacteria) or scanning-dependent (eukaryotes) | Direct 70S/80S binding; multiple factor-independent pathways [49] [17] |
| Initiation Factors | Dependent on multiple factors (eIF2, eIF4F in eukaryotes; IF1, IF2, IF3 in bacteria) | Factor-independent or minimal factor requirement [17] |
| Stress Resistance | Sensitive to stress-induced inactivation of initiation factors | Relatively resistant to various stress conditions [17] |
| Regulatory Potential | High (5' UTR structure, sRNA binding, protein interactions) | Limited (lack 5' UTR regulatory elements) |
| mRNA Stability | Influenced by 5' UTR-mediated mechanisms [13] | Similar half-lives to leadered transcripts with sigA 5' UTR [13] |
Translation Efficiency Measurements Across Systems
Table 2: Quantitative Effects of UTR Features on Translation Efficiency
| UTR Feature | System | Effect on Translation Efficiency | Experimental Method |
|---|---|---|---|
| sigA 5' UTR (123 nt) | M. smegmatis | Decreased apparent translation rate compared to synthetic 5' UTR [13] | Fluorescence reporters, mRNA half-life measurements |
| Leaderless Structure | M. smegmatis | Similar translation efficiency as sigA 5' UTR but lower transcript production rates [13] | Fluorescence reporters, transcript production calculations |
| 3' UTR Length Increase (4-104 nt) | CHO cells | 38-fold increase for poly(A)- mRNA [51] | Luciferase reporter assays |
| C-less 5' UTR Library | E. coli | Highest overall translation efficiency among nucleotide-deficient libraries [50] | sfGFP fluorescence, flow cytometry |
| AU-rich Element Insertion | Human cells | Up to 5-fold increase in protein expression [52] | Luciferase, EGFP, mCherry reporters |
| Alternative 3' UTRs | Drosophila S2 cells | >100-fold variation in repression magnitude [53] | Dual luciferase assays, miRNA targeting |
Experimental Approaches and Methodologies
Reporter Construct Design and Validation
Investigating UTR-mediated translation regulation requires carefully designed reporter systems. The construction of fluorescent protein reporters (e.g., YFP, sfGFP) under control of constitutive promoters enables quantitative assessment of UTR function [13] [50]. For studies in mycobacteria, researchers have successfully employed the pmyc1tetO promoter to drive expression of transcripts containing specific 5' UTRs fused to fluorescent protein coding sequences [13]. Critical validation steps include experimental confirmation of 5' UTR boundaries and translation start codons through mutagenesis approaches, as demonstrated in M. smegmatis where GTG to GTC mutations established the authentic initiation codon [13]. For leaderless transcripts, control constructs with mutated start codons are essential to confirm translation initiation specificity [17].
Dual Luciferase Assay Systems
Dual luciferase reporter systems provide a robust methodology for quantifying translation efficiency and miRNA-mediated repression [53]. This approach typically involves co-transfection of experimental (e.g., Firefly luciferase) and control (e.g., Renilla luciferase) reporters, allowing normalization of translation efficiency against transfection efficiency and cellular variability. The system has been effectively deployed to demonstrate how 3' UTR sequences modulate translatability and miRNA-mediated repression in Drosophila S2 cells, revealing that different 3' UTRs can alter repression magnitude by over 20-fold [53]. These assays can be adapted with various UTR configurations, codon optimization levels, and termination signals (poly(A) tail versus histone stem-loop) to dissect specific regulatory mechanisms [53].
Ribosome Profiling and Computational Modeling
Ribosome profiling (Ribo-seq) has emerged as a powerful technique for genome-wide assessment of translation efficiency by sequencing ribosome-protected mRNA fragments [54] [47]. This approach enables precise measurement of ribosome density at codon resolution, providing insights into both translational efficiency and elongation dynamics. Recent advances integrate ribosome profiling data with absolute quantification of tRNAs, mRNAs, and proteins to derive initiation and elongation rates [47]. State-of-the-art computational tools like RiboNN employ deep convolutional neural networks to predict translation efficiency from mRNA sequence features, capturing how spatial positioning of dinucleotide and trinucleotide features influences translational output [54]. These models demonstrate that the entire mRNA sequence—not just the 5' UTR—jointly determines translation efficiency.
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Reagents and Methods for UTR Translation Research
| Reagent/Method | Function/Application | Example Use Case |
|---|---|---|
| Fluorescent Protein Reporters (YFP, sfGFP) | Quantitative measurement of translation efficiency | M. smegmatis sigA 5' UTR characterization [13] |
| Dual Luciferase Assay Systems | Normalized measurement of translation efficiency and miRNA repression | 3' UTR variant screening in Drosophila S2 cells [53] |
| Ribosome Profiling (Ribo-seq) | Genome-wide mapping of translating ribosomes | Translation efficiency atlas generation across 140+ human cell types [54] |
| Flow Cytometry | High-throughput screening of UTR library variants | Analysis of 5' UTR nucleotide composition libraries in E. coli [50] |
| FLERT (Fleeting mRNA Transfection) | Rapid assessment of mRNA translation in living cells | Leaderless mRNA translation analysis under stress conditions [17] |
| Machine Learning Models (RiboNN) | Prediction of translation efficiency from sequence features | Interpretation of evolutionary constraints in human 5' UTRs [54] |
| Cell-Free Translation Systems | Mechanistic studies of initiation pathways | Factor-independent translation of leaderless mRNAs [49] [17] |
Implications for Therapeutic Development and Synthetic Biology
Understanding UTR-mediated regulation of translation efficiency has profound implications for therapeutic development, particularly in the design of mRNA vaccines and therapeutics. Recent advances demonstrate that engineered AU-rich elements in 3' UTRs can enhance both stability and protein expression by facilitating interactions with RNA-binding proteins like HuR [52]. The sequence "AUUUA" with specific repeats has been shown to increase protein expression up to 5-fold, providing a design principle for therapeutic mRNA optimization [52]. Similarly, the unique properties of leaderless mRNAs—particularly their resistance to cellular stress and reduced dependence on canonical initiation factors—offer potential advantages for therapeutic expression in disease states where standard translation machinery is compromised [17]. In synthetic biology, the systematic analysis of 5' UTR nucleotide composition informs the design of expression cassettes with predictable translation rates, enabling precise tuning of metabolic pathway components [50].
The length and composition of untranslated regions represent fundamental determinants of translation efficiency that operate through conserved yet adaptable mechanisms. The distinction between leadered and leaderless genes exemplifies evolutionary diversification in translational control strategies, with leaderless transcripts employing specialized initiation pathways that provide resilience under stress conditions. Advanced methodologies including reporter assays, ribosome profiling, and machine learning models continue to reveal the complex principles governing UTR-mediated regulation. As research progresses, the integration of quantitative measurements with computational predictions will further elucidate the intricate relationship between mRNA sequence and translational output, enabling more sophisticated engineering of therapeutic mRNAs and synthetic genetic circuits. The ongoing characterization of UTR function across diverse biological systems promises to uncover novel regulatory mechanisms and expand our fundamental understanding of translation control.
The assessment of mRNA stability presents a fundamental challenge in molecular biology, complicated by the intricate crosstalk between transcription and decay processes. This technical guide examines sophisticated methodological approaches required to disentangle these interconnected pathways, with particular emphasis on the distinctions between leadered and leaderless mRNA architectures. The differential stability mechanisms and translational properties of these mRNA classes have profound implications for basic research and therapeutic development, necessitating specialized analytical frameworks for accurate kinetic measurement. By synthesizing recent advances in high-throughput sequencing, computational modeling, and biochemical fractionation, this review provides researchers with a comprehensive toolkit for precise mRNA stability determination across diverse biological contexts and mRNA structural types.
Messenger RNA stability serves as a critical control point in gene regulation, directly influencing the temporal duration and abundance of protein synthesis. The accurate determination of mRNA half-lives is technically challenging due to the constitutive coupling of transcription and decay processes within cells. This interdependence creates a circular gene expression system where mRNA levels represent a steady-state equilibrium between synthesis and degradation rates. For leadered mRNAs—those containing 5' untranslated regions (UTRs)—stability is influenced by multiple structural elements including 5' cap structures, UTR length, upstream open reading frames (uORFs), and nucleotide composition. In contrast, leaderless mRNAs (lmRNAs), which completely lack or possess extremely short 5' UTRs, exhibit distinct regulatory properties and stability determinants that necessitate specialized assessment approaches [55] [17].
The translational initiation pathways employed by leaderless mRNAs differ significantly from their leadered counterparts, contributing to their altered stability profiles. While leadered mRNAs typically utilize cap-dependent scanning mechanisms, leaderless mRNAs can initiate translation through unconventional pathways including direct 80S ribosome binding, eIF2-independent mechanisms, and eIF5B/IF2-assisted initiation [17]. These differences directly impact mRNA decay kinetics, as translation initiation efficiency is intimately connected to degradation pathways. Furthermore, technical challenges in stability assessment are compounded by the discovery that transcriptional start site heterogeneity generates multiple transcript isoforms from single genes, each potentially exhibiting distinct stability characteristics [56]. This technical overview addresses these complexities by providing a detailed framework for discriminating transcription and decay contributions to mRNA abundance across diverse mRNA architectures.
Fundamental Differences Between Leadered and Leaderless mRNAs
Structural and Functional Characteristics
Table 1: Comparative Analysis of Leadered and Leaderless mRNA Architectures
| Feature | Leadered mRNAs | Leaderless mRNAs |
|---|---|---|
| 5' UTR Presence | Present (typically 20-200 nt) | Absent or very short (0-5 nt) |
| Translation Initiation | Canonical cap-dependent scanning | Multiple unconventional pathways |
| Initiation Factor Requirement | eIF2, eIF4F dependent | eIF2/eIF4F independent under stress |
| Ribosome Recruitment | 40S-mediated with scanning | Direct 80S binding or specialized mechanisms |
| Stability Determinants | 5' cap, 5' UTR elements, poly(A) tail | 5' terminal structure, ribosome protection |
| Prevalence | Majority of eukaryotic transcripts | Varies by organism (1-70% of transcriptome) |
| Stress Resistance | Sensitive to eIF2 phosphorylation | Relatively resistant to various stresses |
The structural dichotomy between leadered and leaderless mRNAs extends beyond mere presence or absence of a 5' UTR to encompass fundamental differences in regulatory capacity and protein output control. Leadered mRNAs contain complex regulatory information within their 5' UTRs, including binding sites for RNA-binding proteins, upstream AUG codons (uAUGs), and secondary structure elements that profoundly influence translation efficiency and mRNA stability [56]. For example, uAUGs in leadered transcripts are associated with reduced translation efficiency and targeting for nonsense-mediated mRNA decay (NMD), effectively coupling translational regulation with decay pathways [56].
Leaderless mRNAs represent molecular relics of ancient translation initiation pathways yet remain functionally significant across diverse taxa. These transcripts are particularly abundant in Archaea and Actinobacteria, with Haloferax volcanii exhibiting approximately 72% leaderless transcripts and Mycobacterium tuberculosis containing approximately 22% [55]. The phylogenetic distribution suggests lmRNAs may represent ancestral mRNA forms, with their persistence in modern organisms indicating specialized functional roles. In eukaryotes, nuclear-encoded leaderless transcripts are widely represented across primitive unicellular organisms and demonstrate unique translational properties including stress resistance to mTOR inhibition and oxidative stress [17]. This resilience likely stems from their ability to utilize multiple initiation pathways when canonical mechanisms are compromised.
Methodological Approaches for Disentangling Transcription and Decay
High-Throughput Sequencing-Based Platforms
PERSIST-seq: Parallel Evaluation of mRNA Stability and Translation
The PERSIST-seq (Pooled Evaluation of mRNA in-solution Stability, and In-cell Stability and Translation RNA-seq) platform enables systematic determination of how UTR sequences, coding sequences, and RNA structural elements influence mRNA translation and stability parameters simultaneously [57]. This approach utilizes a combinatorial library design with barcoded mRNA variants to facilitate parallel assessment of multiple stability metrics.
Experimental Workflow:
- Library Construction: A diverse library of mRNA sequences is synthesized with variations in 5' UTR, CDS, and 3' UTR regions. Each template includes a shared T7 promoter, unique barcodes in the 3' UTR, and constant regions for pooled amplification.
- In Vitro Transcription: The DNA library is transcribed in vitro with simultaneous 5' capping and 3' polyadenylation to generate mature mRNA molecules.
- Cellular Transfection: The pooled mRNA library is introduced into cells for in-cell stability assessment or maintained in solution for in-solution stability measurements.
- Polysome Profiling: Transfected cells are treated with cycloheximide to arrest translating ribosomes, followed by sucrose density fractionation to separate mRNAs based on ribosome occupancy.
- Sequencing and Analysis: Barcode sequencing of input and fractionated samples enables quantification of ribosome load (translation efficiency) and mRNA abundance over time (stability).
PERSIST-seq analysis revealed that in-cell stability is a greater determinant of protein output than high ribosome load, challenging conventional assumptions that translation efficiency primarily governs protein production [57]. Furthermore, this approach demonstrated that highly structured "superfolder" mRNAs can be designed to improve both stability and expression, particularly when combined with pseudouridine nucleoside modification.
TL-seq and TATL-seq: Transcript Leader Analysis
TL-seq (Transcript Leader sequencing) combines enzymatic capture of m7G-capped mRNA 5' ends with high-throughput sequencing to map transcript leader boundaries genome-wide [56]. The related TATL-seq (Translation-Associated Transcript Leader sequencing) integrates TL-seq with polysome fractionation to simultaneously annotate TLs and assess their translational function.
Experimental Workflow:
- RNA Processing: Fragmented and size-selected RNA is treated with phosphatase to reduce non-capped 5' ends to hydroxyl groups.
- Cap Conversion: Treatment with pyrophosphatase cleaves two phosphates from the cap structure, yielding 5' monophosphorylated RNA.
- Adapter Ligation: RNA ligase selectively ligates adapters to the 5' monophosphate (formerly capped) fragments.
- Library Preparation: Ligated RNA fragments are converted to DNA sequencing libraries using standard protocols.
- Sequencing and Peak Calling: High-throughput sequencing identifies transcription start sites, with computational peak-calling distinguishing authentic TSSs from background.
TL-seq applications have revealed surprising transcriptional heterogeneity, including the discovery that 6% of protein-coding genes in yeast contain transcription initiation sites within their coding regions, concentrated near 5' ends of ORFs [56]. These internal start sites produce truncated mRNAs that are actively translated, contributing to proteome diversity and complicating stability assessments.
Computational Modeling of mRNA Dynamics
Mathematical modeling approaches provide powerful tools for quantifying the individual contributions of transcription and decay to mRNA abundance. These models typically treat mRNA levels as dynamic systems where changes in abundance reflect the balance between synthesis and degradation.
Basic Kinetic Model:
Where ksynthesis represents the transcription rate and kdecay the first-order decay constant.
For leadered and leaderless mRNAs, different factors influence these rate constants. Leadered mRNA transcription rates are influenced by promoter elements and transcription factor binding, while decay rates are affected by 5' UTR features, coding sequence elements, and 3' UTR determinants. Leaderless mRNA kinetics may be influenced by different factors, including direct ribosome interactions and specialized degradation pathways.
Advanced modeling approaches incorporate crosstalk factors that coordinately regulate both transcription and decay. For example, RNA-binding proteins like Sfp1 and Puf3 in yeast can influence both mRNA synthesis and degradation rates, creating buffering or enhancing effects on steady-state mRNA levels [58]. When transcription and mRNA degradation act at compensatory rates, mRNA buffering occurs, maintaining approximately constant levels despite regulatory changes. Conversely, when both processes act additively, enhanced gene expression regulation occurs [58].
Diagram Title: mRNA Lifecycle: Transcription-Decay Crosstalk
Advanced Analytical Techniques for mRNA Characterization
Electrophoretic and Chromatographic Methods
Capillary gel electrophoresis (CGE) and ion-pair reversed-phase liquid chromatography (IP-RP LC) provide high-resolution separation of mRNA species based on size and hydrophobicity, respectively. These techniques are essential for assessing mRNA integrity and identifying degradation products [59]. CGE separates mRNA molecules based on their size-to-charge ratio in a narrow capillary filled with conductive buffer, enabling precise quantification of full-length and truncated species. For therapeutic mRNA applications, regulatory agencies require demonstration of >55% intact mRNA in final products, highlighting the importance of rigorous integrity assessment [59].
Size exclusion chromatography (SEC) complements these approaches by identifying mRNA aggregates based on size separation. When combined with multi-angle light scattering detection, SEC provides absolute molecular weight determinations that verify mRNA integrity and detect aberrant multimerization states that may impact stability and function.
Mass Spectrometric and Sequencing Approaches
Liquid chromatography-tandem mass spectrometry (LC-MS/MS) enables detailed characterization of mRNA chemical modifications and sequence verification through oligonucleotide mapping [59]. This approach is particularly valuable for quantifying nucleoside modifications such as pseudouridine (Ψ) and N1-methyl pseudouridine (m1Ψ), which significantly impact mRNA immunogenicity, stability, and translation efficiency [60].
Direct RNA sequencing using platforms from Oxford Nanopore Technologies allows full-length mRNA characterization without reverse transcription, preserving modification information that would be lost in cDNA-based methods. This approach can detect chemical modifications and poly(A) tail length heterogeneity simultaneously, providing comprehensive mRNA characterization [59].
Table 2: Analytical Techniques for mRNA Stability Assessment
| Technique | Application | Key Metrics | Considerations for Leadered vs. Leaderless mRNAs |
|---|---|---|---|
| PERSIST-seq | Simultaneous measurement of in-cell stability, translation efficiency, and in-solution stability | Ribosome load, degradation kinetics | Leaderless mRNAs may show different structure-stability relationships |
| TL-seq/TATL-seq | Genome-wide mapping of transcript leaders and association with translation | TSS identification, TL heterogeneity | Leaderless mRNAs lack traditional TLs; internal TSSs may be significant |
| CGE | Integrity assessment and size-based separation | Full-length percentage, size distribution | Leaderless mRNAs typically shorter; require different size standards |
| IP-RP LC | Separation based on hydrophobicity | Purity, impurity profiling | Different nucleotide composition may alter retention behavior |
| LC-MS/MS | Modification characterization and sequence verification | Modification quantification, sequence confirmation | Leaderless mRNAs may have different modification preferences |
| Direct RNA Sequencing | Full-length sequence and modification analysis | Modification detection, poly(A) tail length | Direct 5' end sequencing crucial for leaderless mRNA validation |
The Researcher's Toolkit: Essential Reagents and Methodologies
Table 3: Essential Research Reagents for mRNA Stability Studies
| Reagent/Method | Function | Application Notes |
|---|---|---|
| PERSIST-seq Library | Parallel assessment of mRNA stability and translation | Enables high-throughput screening of sequence-stability relationships |
| TL-seq Enzymatic Kit | Specific capture of m7G-capped mRNA 5' ends | Critical for accurate TSS mapping and TL heterogeneity assessment |
| Polysome Profiling Reagents | Separation of mRNAs based on ribosome occupancy | Requires cycloheximide treatment to arrest translation |
| Metabolic Labeling Agents | Temporal tracking of newly synthesized mRNA | 4-thiouridine (4sU) and analogs enable precise pulse-chase experiments |
| In-line Hydrolysis Buffer | Assessment of intrinsic RNA stability | Controlled pH and temperature conditions essential for reproducibility |
| Pseudouridine Modified NTPs | Enhanced stability and reduced immunogenicity | Impacts both stability and translational properties; concentration optimization required |
| Cap Analogs | 5' end modification for translation enhancement | ARCA and clean cap analogs improve capping efficiency and expression |
| Poly(A) Tail Length Standards | Calibration for tail length assessment | Essential for accurate poly(A) tail measurements by sequencing or electrophoresis |
Special Considerations for Leadered vs. Leaderless mRNA Analysis
The analytical strategies for assessing mRNA stability must be adapted based on the fundamental structural and functional differences between leadered and leaderless mRNAs. For leadered mRNAs, particular attention must be paid to 5' UTR elements that significantly influence stability, including upstream AUGs (uAUGs), secondary structures, and protein-binding motifs. Research has demonstrated that uAUGs are associated with reduced translation efficiency and targeting for nonsense-mediated mRNA decay, creating a direct link between translation initiation and mRNA stability [56]. Additionally, short TLs are associated with inefficient translation initiation at annotated start codons and increased initiation at downstream AUGs, frequently resulting in out-of-frame translation and subsequent NMD [56].
For leaderless mRNAs, alternative analytical approaches are required due to their unique properties. These mRNAs often exhibit non-canonical initiation pathways that can render their translation resistant to conditions that inhibit canonical initiation, such as eIF2 phosphorylation during stress responses [17]. This resilience necessitates specialized stress protocols to fully characterize their stability properties. Furthermore, the absence of a 5' UTR eliminates many regulatory elements that influence leadered mRNA stability, requiring focus on alternative stability determinants such as 5' terminal nucleotides and ribosome protection effects.
Diagram Title: Translation Initiation Pathways Under Stress
The accurate assessment of mRNA stability requires sophisticated methodological approaches that simultaneously quantify transcription and decay parameters while accounting for the fundamental differences between leadered and leaderless mRNA architectures. The integration of high-throughput sequencing platforms like PERSIST-seq and TL-seq with advanced computational models provides a powerful framework for dissecting these complex relationships. Future methodological developments will likely focus on single-molecule imaging approaches that visualize transcription and decay events in real-time, further refining our understanding of mRNA dynamics across diverse biological contexts.
For therapeutic applications, the stability principles governing leadered and leaderless mRNAs present distinct optimization opportunities. Leadered mRNAs benefit from UTR engineering strategies that enhance stability and translation, such as incorporating AU-rich elements that promote HuR binding [52]. In contrast, leaderless mRNAs offer advantages in stress-resistant expression but present unique design challenges due to their simplified architecture. As mRNA therapeutics expand beyond vaccination to protein replacement and gene editing applications, the nuanced understanding of stability determinants for both mRNA classes will be essential for designing next-generation therapeutics with optimized pharmacokinetics and tissue-specific expression profiles.
For decades, Escherichia coli has served as the primary model organism for deciphering fundamental bacterial processes, including the canonical mechanisms of translation initiation. The established paradigm, derived predominantly from E. coli studies, describes a process where the 30S ribosomal subunit binds to a Shine-Dalgarno (SD) sequence within the 5' untranslated region (5' UTR) of an mRNA, facilitating positioning at the start codon [61] [14]. This leadered initiation mechanism has long been considered the standard for bacterial translation. However, the overreliance on E. coli as a universal model has created a significant knowledge gap, particularly evident when studying bacterial pathogens that deviate from this canonical pattern. A critical limitation arises when translating findings related to gene regulation and protein synthesis from E. coli to pathogenic species like Mycobacterium tuberculosis, the causative agent of tuberculosis. Research has revealed that nearly one-quarter of transcripts in M. tuberculosis and the model organism Mycobacterium smegmatis are leaderless, meaning they completely lack a 5' UTR and the associated SD sequence [61] [62]. This prevalence stands in stark contrast to E. coli, where leaderless transcripts are rare and often associated with mobile genetic elements [61] [14]. This discrepancy underscores a fundamental challenge in molecular bacteriology: mechanisms elucidated in a model organism may not fully represent the biological reality of distantly related pathogens, potentially hampering drug development efforts that target gene expression.
The implications of these differences extend beyond basic science to therapeutic design. Understanding how pathogens like M. tuberculosis regulate gene expression is essential for developing interventions that disrupt its survival within the human host [13]. If the translational landscape of a pathogen differs significantly from the E. coli model, strategies designed to inhibit protein synthesis based on that model may prove ineffective. This technical guide examines the key differences between leadered and leaderless translation systems, provides methodologies for cross-species experimental validation, and outlines a framework for effectively translating molecular findings from model organisms to bacterial pathogens.
Fundamental Distinctions: Leadered vs. Leaderless Genes
Mechanistic and Sequence Requirements
Leaderless and leadered genes represent two distinct paradigms of translation initiation with different mechanistic requirements. The table below summarizes the core features that differentiate these two types of gene structures.
Table 1: Key Characteristics of Leadered and Leaderless Translation Initiation
| Feature | Leadered Translation | Leaderless Translation |
|---|---|---|
| 5' UTR | Present (median ~40-56 nt in mycobacteria) [13] | Absent [61] |
| Shine-Dalgarno Sequence | Required for robust initiation [61] [14] | Absent [61] |
| Initiating Ribosome | 30S ribosomal subunit [61] [14] | 70S ribosome (full) [61] [14] |
| Start Codon Requirement | ATG, GTG, TTG, ATT (in mycobacteria) [61] | 5' ATG or GTG is necessary and sufficient in mycobacteria [61] |
| Initiation Factor Sensitivity | Stimulated by IF3 [61] [14] | Inhibited by IF3; stimulated by IF2 [61] [14] |
For leadered transcripts, the SD sequence within the 5' UTR is critical for recruitment and positioning of the 30S ribosomal subunit. In mycobacteria, leadered initiation can use alternative start codons (ATG, GTG, TTG, and ATT), demonstrating a flexibility that must be accounted for when annotating genes in pathogens [61]. In contrast, leaderless translation requires a fundamentally different mechanism. The absence of a 5' UTR means that fully assembled 70S ribosomes must bind directly to the 5' end of the mRNA, where the start codon itself serves as the primary recognition signal [61] [12]. Experimental validation in mycobacteria has demonstrated that an ATG or GTG at the 5' terminal position is both necessary and sufficient for this process [61]. Furthermore, leaderless translation is differentially affected by initiation factors; it is inhibited by initiation factor 3 (IF3), which typically acts to stabilize 30S subunit binding to SD sequences, while being stimulated by initiation factor 2 (IF2) [61] [14].
Functional and Evolutionary Distribution
The distribution of leaderless genes across the bacterial kingdom is not random but reveals significant evolutionary and functional patterns.
Table 2: Distribution and Functional Associations of Leaderless Genes
| Bacterial Group | Prevalence of Leaderless Genes | Notable Functional Associations |
|---|---|---|
| E. coli | Rare (1.2-3%); often phage-derived [61] [63] | Mobile genetic elements [61] [14] |
| Mycobacterium tuberculosis | High (~25-26% of transcripts) [61] [62] | Stress response, toxin-antitoxin systems, non-replicating persistence [62] [63] |
| Actinobacteria | High (>20% of genes) [10] | Not specified in search results |
| Archaea | Highly prevalent [10] [61] | Proposed ancient, ancestral mechanism [10] [61] |
Computational analysis of 953 bacterial genomes reveals that leaderless genes are widespread, though not dominant, across many bacterial groups [10]. They are particularly abundant in Actinobacteria (the phylum containing Mycobacterium) and Deinococcus-Thermus, where they can constitute over twenty percent of all genes [10]. Evolutionary analyses suggest that the proportion of leaderless genes in bacteria has a decreasing trend over time, supporting the hypothesis that leaderless initiation may be an ancient mechanism used by the last universal common ancestor (LUCA) [10] [61]. This theory is bolstered by the prevalence of leaderless genes in archaea and the observation that leaderless transcripts can be translated in all three domains of life [10].
Critically, in pathogens like M. tuberculosis, leaderless genes are not randomly distributed but are functionally enriched. They are markedly associated with stress adaptation, including toxin-antitoxin modules and functions important during growth arrest [62] [63]. This functional specialization has direct implications for pathogenesis, as non-replicating persistence is a key feature of chronic tuberculosis infection.
Experimental Approaches: Validating Translation Mechanisms in Pathogens
Genome-Wide Mapping Techniques
Overcoming model organism limitations requires empirical validation in the pathogen of interest. The following experimental workflows allow for genome-wide mapping of transcriptional and translational events without relying on E. coli-derived assumptions.
Diagram 1: Genome-wide Transcriptional Start Site (TSS) Mapping
The first critical step is the precise mapping of transcriptional start sites (TSSs). As illustrated in Diagram 1, this process begins with bacterial culture under relevant conditions, followed by RNA extraction. A key step is the enrichment of 5' triphosphate transcripts, which distinguishes primary transcripts from processed RNAs [62]. Subsequent RNA sequencing and bioinformatic analysis identifies TSSs genome-wide. A transcript is defined as leaderless if its TSS overlaps exactly with the annotated start codon, indicating the absence of a 5' UTR [62] [12].
Diagram 2: Ribosome Profiling to Map Translation Initiation
To directly monitor translation, ribosome profiling (Ribo-seq) is employed. This technique, outlined in Diagram 2, involves treating bacterial cultures with a ribosome-stalling agent, followed by nuclease digestion that degrades mRNA regions not protected by bound ribosomes. The protected fragments ("ribosome footprints") are then purified and sequenced [61]. The 5' boundaries of these footprints indicate the exact positions where ribosomes initiate translation. For leaderless transcripts, the 5' boundaries of RNA-seq reads (indicating transcription start) and ribosome profiling reads (indicating translation start) coincide at the start codon [61] [12].
Targeted Reporter Assays for Functional Validation
While genome-wide methods identify potential leaderless transcripts, functional validation requires controlled reporter assays. The following protocol describes a methodology for comparing leadered versus leaderless translation, particularly during stress conditions relevant to pathogenesis.
Table 3: Key Research Reagents for Leaderless Translation Studies
| Reagent/Tool | Function/Description | Application Example |
|---|---|---|
| Integrating Reporter Vector (e.g., pMV306) | Stable, single-copy chromosomal integration | Ensures consistent gene dosage for comparing expression between constructs [63] |
| Transcriptional Terminators (e.g., trp terminator) | Prevents read-through transcription from plasmid | Isolates the effect of the cloned 5' regulatory sequences [63] |
| Firefly Luciferase (ffluc) Reporter | Sensitive, quantifiable enzymatic output | Allows precise measurement of translation efficiency over time and under stress [63] |
| Candidate Promoter/5' Regions | Genomic regions fused to reporter | Compares native leadered (e.g., desA1) vs. leaderless (e.g., desA2) structures [63] |
Experimental Protocol: Stress-Resilience Translation Assay
Construct Generation:
- Clone the promoter and 5' region (including UTR for leadered, or start codon only for leaderless) from a pathogen gene of interest (e.g., M. tuberculosis desA1 for leadered, desA2 for leaderless) upstream of a promoter-less firefly luciferase (ffluc) gene in an integrating vector [63].
- Ensure the leaderless construct is generated such that the start codon is the first nucleotide at the 5' end of the transcript.
- Introduce constructs into the target pathogen (e.g., M. tuberculosis) to generate isogenic reporter strains.
Baseline Measurement:
- Grow reporter strains under optimal, exponential-phase conditions.
- Harvest samples and measure luminescence as a direct indicator of translation output. Normalize to cell density or protein content.
- Under these conditions, leaderless translation in mycobacteria is robust and can be comparable to leadered translation [63].
Stress Challenge:
- Subject the reporter strains to relevant stress conditions that mimic the host environment. For M. tuberculosis, this includes:
- Nutrient Starvation: Transfer to minimal media without a carbon/nitrogen source.
- Nitric Oxide Exposure: Treat with a nitric oxide donor at a sub-lethal concentration.
- Macrophage Infection: Infect murine or human macrophage cell lines and monitor over time [63].
- Collect samples at multiple time points for luminescence measurement and RNA extraction.
- Subject the reporter strains to relevant stress conditions that mimic the host environment. For M. tuberculosis, this includes:
Data Analysis:
- Quantify changes in luminescence for leadered vs. leaderless reporters during stress.
- Extract RNA and perform RT-qPCR on the reporter transcript to control for transcription-level effects.
- Calculate the translation efficiency (Luminescence / mRNA level) for each construct.
- Expected Outcome: Studies show that leaderless translation in M. tuberculosis remains more stable than leadered translation during adaptation to nutrient starvation, nitric oxide exposure, and macrophage infection, indicating a resilience advantage under stress [63].
A Path Forward: Strategic Framework for Cross-Species Translation
To systematically address the limitations of model organisms, researchers should adopt a structured framework that prioritizes empirical validation in pathogens. The following strategic recommendations are derived from the collective findings:
Profile Before Assuming: Begin research on a pathogen's gene regulation by empirically defining its transcription and translation landscape using TSS mapping [62] and ribosome profiling [61]. Do not assume the E. coli paradigm applies.
Validate Functionally: Use controlled reporter assays, as described in Section 3.2, to test the translation efficiency and regulatory properties of specific leadered and leaderless genes under physiologically relevant conditions [63].
Contextualize Biologically: Interpret the function of leaderless genes within the pathogen's specific life cycle. For M. tuberculosis, the association of leaderless transcripts with stress response and toxin-antitoxin modules suggests they are key players in persistence [62] [63], making them potential targets for disrupting latent infection.
Account for Technical Artefacts: Be aware that standard RNA-seq sample preparation, which often includes ribodepletion, may inadvertently deplete leaderless transcripts, as their start codon can base-pair with the anti-SD sequence on the 16S rRNA used in ribodepletion protocols. Use poly(A) tailing methods or 5' triphosphate enrichment to avoid this bias [62].
Explore Mechanistic Differences: Investigate pathogen-specific translational machinery. For example, the distinct sequence preference of mycobacterial RNase E (cleavage upstream of cytidines) compared to the E. coli enzyme influences mRNA degradation rates and can differentially affect leadered and leaderless transcripts [64].
By integrating these approaches, the scientific community can move beyond the constraints of the E. coli model and develop a more accurate, pathogen-centric understanding of bacterial gene regulation, ultimately accelerating the discovery of novel therapeutic targets.
In the field of synthetic biology, the precise control of gene expression is fundamental to engineering predictable and efficient biological systems. While promoters and coding sequences have traditionally received significant attention, untranslated regions (UTRs) have emerged as equally critical components for fine-tuning gene expression. UTRs are the non-coding sections of messenger RNA (mRNA) that flank the protein-coding sequence; the 5' UTR is located upstream of the start codon, while the 3' UTR is found downstream of the stop codon [65]. These regions serve as central hubs for post-transcriptional regulation, influencing mRNA stability, localization, and translation efficiency [66] [5]. In bacteria, the 5' UTR typically contains the Shine-Dalgarno (SD) sequence, which facilitates ribosome binding and translation initiation [1]. However, an important distinction exists in prokaryotic systems between "leadered" genes, which possess a 5' UTR, and "leaderless" genes, which completely lack a 5' UTR and initiate translation directly at the start codon [13] [10].
The selection of appropriate UTRs is not merely a technical consideration but a fundamental aspect of synthetic biology design that can determine the success of genetic constructs. Research has revealed that bacterial species exhibit remarkable diversity in their use of leadered versus leaderless genes. For instance, in Mycobacterium tuberculosis, approximately 25-26% of genes are leaderless, a significantly higher proportion than observed in model organisms like Escherichia coli [62]. This distribution has profound implications for designing expression systems optimized for specific bacterial chassis. Furthermore, the length and composition of 5' UTRs in leadered genes vary substantially, with median lengths of 48 and 56 nucleotides in M. smegmatis and M. tuberculosis, respectively [13]. This biological diversity provides a rich toolkit for synthetic biologists but also necessitates a systematic approach to UTR selection based on well-defined design principles and empirical data, which this review will explore in depth.
Biological Foundations: Leadered vs. Leaderless Gene Architectures
Molecular Mechanisms of Translation Initiation
The fundamental distinction between leadered and leaderless genes lies in their mechanisms of translation initiation. For leadered transcripts, the process begins with the binding of the ribosomal complex to the Shine-Dalgarno (SD) sequence within the 5' UTR, typically located 3-10 nucleotides upstream of the initiation codon [10] [1]. This SD sequence (5'-AGGAGGU-3') base-pairs with the complementary anti-SD sequence at the 3' end of the 16S rRNA, positioning the ribosome correctly to initiate translation at the downstream start codon. This mechanism allows for additional regulatory elements within the 5' UTR to influence translation efficiency, including upstream open reading frames (uORFs), RNA secondary structures, and binding sites for proteins or small RNAs [13].
In contrast, leaderless transcripts completely lack 5' UTRs and therefore initiate translation directly at the start codon, which is exposed at the very 5' end of the mRNA [10]. This initiation mechanism bears similarity to eukaryotic translation and is thought to represent an evolutionarily ancient process [10]. Early research suggested that leaderless genes might be translated less efficiently in model organisms like E. coli [13], but studies in mycobacteria have demonstrated that leaderless transcripts can be translated robustly, indicating that translation efficiency is organism-dependent and influenced by cellular context [13].
Distribution Across Bacterial Species and Functional Associations
The prevalence of leaderless genes varies dramatically across bacterial species, suggesting distinct evolutionary adaptations. Genomic analyses of 953 bacterial and 72 archaeal genomes reveal that leaderless genes are "widespread, although not dominant, in a variety of bacteria" [10]. Certain bacterial groups show particularly high proportions of leaderless genes, with Actinobacteria and Deinococcus-Thermus exhibiting more than 20% leaderless genes in their genomes [10]. In M. tuberculosis, this proportion reaches approximately 26% of all genes [62].
Functionally, leadered and leaderless gene architectures are associated with different biological processes. In M. tuberculosis, genes encoding proteins with active growth functions are "markedly depleted from the leaderless transcriptome" [62]. Instead, leaderless genes show significant enrichment in stress response pathways and toxin-antitoxin modules [62]. Furthermore, research has demonstrated that the "abundance of leaderless mRNAs increases during starvation-induced growth arrest" [62], suggesting that the leaderless architecture may represent an adaptive strategy for maintaining essential gene expression under nutrient limitation. This functional specialization has important implications for synthetic biology applications, particularly when designing expression systems for stress-resistant industrial organisms or persistent pathogens.
Quantitative Comparison of Leadered and Leaderless Systems
Performance Metrics for Gene Expression
Understanding the quantitative performance differences between leadered and leaderless architectures is essential for informed UTR selection in synthetic biology. Research using fluorescence reporters in Mycobacterium smegmatis has revealed that these two systems differ across multiple parameters of gene expression, including transcript production rates, mRNA half-life, and translation efficiency [13]. The table below summarizes key comparative metrics derived from experimental studies:
Table 1: Quantitative comparison of leadered and leaderless gene expression characteristics
| Parameter | Leadered Genes | Leaderless Genes | Experimental System |
|---|---|---|---|
| 5' UTR Length | Median: 48 nt (M. smegmatis), 56 nt (M. tuberculosis) [13] | 0 nt (by definition) [10] | Genome-wide analysis |
| Transcript Production Rate | Variable; sigA 5' UTR showed increased rate [13] | Lower predicted rates [13] | Fluorescence reporters in M. smegmatis |
| mRNA Half-Life | Variable; sigA 5' UTR conferred shorter half-life [13] | Similar to sigA 5' UTR [13] | Fluorescence reporters in M. smegmatis |
| Translation Efficiency | Variable; sigA 5' UTR decreased efficiency [13] | Similar to sigA 5' UTR [13] | Fluorescence reporters in M. smegmatis |
| Protein/MRNA Ratio | No systematic difference detected [13] | No systematic difference detected [13] | Global comparison in M. tuberculosis |
| Prevalence in Bacterial Genomes | Majority of genes in most bacteria [10] | 20%+ in Actinobacteria, 26% in M. tuberculosis [10] [62] | Genomic analysis of 953 bacteria |
Impact of 5' UTR Features on Leadered Gene Expression
For leadered genes, specific features of the 5' UTR significantly influence expression outcomes. Research has demonstrated that the length and sequence composition of 5' UTRs can dramatically alter both mRNA stability and translation efficiency [13]. For instance, the long 5' UTR of the sigA gene (123 nt in M. smegmatis) was found to confer an "increased transcript production rate, shorter mRNA half-life, and decreased apparent translation rate compared to a synthetic 5' UTR commonly used in mycobacterial expression plasmids" [13]. This illustrates how native 5' UTRs can possess complex regulatory properties that might be undesirable for standard expression systems.
Secondary structure formation within 5' UTRs plays a particularly important role in regulating transcript stability and translation. Stable secondary structures can protect against 5' scanning by ribonucleases, thereby increasing mRNA half-life [13]. However, these same structures can potentially impede ribosome scanning and binding, thereby reducing translation efficiency [13] [1]. This trade-off creates a design challenge for synthetic biologists seeking to optimize expression levels. Additionally, 5' UTRs can contain binding sites for regulatory proteins and small RNAs that further modulate gene expression in response to cellular conditions [13]. Understanding these complex interactions is essential for predicting the behavior of synthetic genetic constructs.
UTR Design Strategies for Synthetic Biology Applications
Engineering 5' Regulatory Sequences
Synthetic biology has developed multiple methodologies for creating and optimizing 5' regulatory sequences (RES), which encompass both promoters and 5' UTRs [67]. These approaches can be categorized into four main strategies, each with distinct advantages and applications:
Table 2: Engineering strategies for 5' regulatory sequences in synthetic biology
| Strategy | Methodology | Key Features | Examples/Applications |
|---|---|---|---|
| Hybrid RES | Combination of known DNA parts through shuffling or recombination [67] | Generates combinatorial libraries; leverages existing characterized parts | tac promoter (trp + lacUV5) [67] |
| Mutated RES | Introduction of random mutations via error-prone PCR [67] | Creates variants with a range of activities; no prior knowledge required | Varying strength of constitutive promoters [67] |
| Semi-artificial RES | Known core motifs combined with random flanking nucleotides [67] | Balances rational design with exploration of novel sequence space | Saturation mutagenesis around -10/-35 boxes [67] |
| Artificial RES | Completely random nucleotide sequences [67] | Maximum novelty; valuable for non-model organisms | De novo RES generation for novel hosts [67] |
The selection of an appropriate engineering strategy depends on multiple factors, including the host organism, desired expression characteristics, and available screening capacity. For model organisms with well-characterized parts libraries, hybrid approaches often provide the most predictable outcomes. However, when working with non-model organisms or seeking novel regulatory functions, artificial or semi-artificial approaches may be more productive despite requiring more extensive screening.
Design Considerations for Leaderless Constructs
The design of leaderless constructs presents unique considerations compared to traditional leadered approaches. Since leaderless mRNAs completely lack 5' UTRs, the nucleotide context surrounding the start codon becomes critically important. Research suggests that a "requirement seems to be a lack of secondary structure near the initiation codon" for efficient translation of leaderless transcripts [1]. This contrasts with leadered genes, where moderate secondary structure can sometimes enhance stability without completely blocking translation.
Start codon selection also represents an important design parameter. While AUG is the most common initiation codon, both GUG and UUG can serve as alternative start codons for leaderless genes, albeit typically with reduced efficiency [13]. The immediate downstream sequence following the start codon can also influence translation initiation rates, as these nucleotides may affect ribosome binding or stability. When designing synthetic leaderless constructs, it is often advisable to include the natural coding sequence context from efficiently expressed native leaderless genes, as this may contain optimized sequence features that have evolved for robust translation.
Experimental Approaches for UTR Characterization and Validation
Methodologies for Functional Analysis
Rigorous characterization of UTR function is essential for developing reliable synthetic biology systems. The following experimental workflow provides a comprehensive approach for evaluating UTR performance:
This integrated approach enables researchers to dissect UTR function at multiple regulatory levels. As demonstrated in recent studies, combining fluorescence reporters with measurements of "protein abundance, mRNA abundance, and mRNA half-life" allows researchers to "calculate relative transcript production rates" and identify the specific step in gene expression most affected by UTR sequence [13]. This multi-level analysis is particularly important because UTRs can influence both transcriptional and post-transcriptional processes.
For specialized applications, more advanced methodologies may be employed. Ribosome profiling (Ribo-seq) provides genome-wide information about translated regions, including upstream open reading frames (uORFs) that might initiate at non-canonical start codons [5]. Massively parallel reporter assays (MPRAs) enable high-throughput screening of thousands of UTR variants simultaneously, generating comprehensive datasets that link sequence to function [68]. These approaches are particularly valuable for building predictive models of UTR activity and identifying novel regulatory elements.
Essential Research Reagents and Tools
The experimental characterization of UTRs relies on specialized reagents and methodologies. The following table catalogues key solutions employed in this research domain:
Table 3: Research reagent solutions for UTR characterization
| Category | Specific Reagents/Methods | Function/Application | Examples from Literature |
|---|---|---|---|
| Reporter Systems | Fluorescent proteins (YFP), epitope tags (6×His) | Quantitative measurement of gene expression | YFP with C-terminal 6×His tag in M. smegmatis [13] |
| Expression Vectors | Constitutive promoters (pmyc1tetO), inducible systems | Controlled expression of test constructs | pmyc1tetO promoter for constitutive expression [13] |
| Analytical Tools | qRT-PCR, RNA-seq, ribosome profiling | Multi-level analysis of gene expression | mRNA half-life measurements [13] |
| Library Technologies | Error-prone PCR, oligonucleotide synthesis, MPRAs | Generation and screening of UTR variant libraries | Massively parallel reporter assays [68] |
| Bioinformatics Tools | Sequence analysis, folding algorithms, motif discovery | In silico prediction of UTR function | Secondary structure prediction [13] |
These research tools enable the comprehensive functional analysis necessary for rational UTR design. Fluorescent reporter systems, particularly when paired with different promoter systems, allow for rapid screening of UTR libraries under various growth conditions [13] [67]. The combination of experimental data with bioinformatic predictions creates a powerful framework for understanding sequence-function relationships in UTRs.
Applications in Therapeutic Development and Industrial Biotechnology
UTR Engineering for Bacterial Pathogen Research
UTR optimization has significant implications for research on bacterial pathogens, particularly for understanding persistence and virulence mechanisms. In M. tuberculosis, the unusually high prevalence of leaderless genes (26%) appears to represent an adaptive strategy for "nonreplicating persistence" within the host [62]. The observation that the "overall representation of leaderless mRNAs increases during starvation-induced growth arrest" [62] suggests that synthetic biology approaches targeting these transcripts could potentially disrupt bacterial persistence. By engineering reporter constructs with native M. tuberculosis UTRs, researchers can monitor bacterial metabolic states and identify conditions that trigger persistence-related gene expression patterns.
UTR engineering also facilitates the development of diagnostic tools and antibacterial screening platforms. Synthetic genetic circuits incorporating stress-responsive UTRs can be designed to report on antibiotic efficacy or identify compounds that specifically target persistent bacteria. For instance, promoters and UTRs from leaderless genes that are upregulated during starvation could drive expression of fluorescent reporters in bacterial screening assays, enabling the identification of compounds that remain effective against non-replicating populations. These applications demonstrate how understanding native UTR function can inform both basic research and therapeutic development for challenging bacterial pathogens.
Optimization of Industrial Expression Systems
In industrial biotechnology, UTR selection plays a crucial role in optimizing product yields and cellular fitness. The finding that "genes intolerant to loss of function have longer and more complex 5' UTRs" [5] suggests that native regulatory mechanisms employ UTR complexity to maintain precise expression of dosage-sensitive genes. Synthetic biologists can leverage this principle when expressing heterologous enzymes or biosynthetic pathways that may place metabolic burdens on host cells. By incorporating appropriately complex UTRs, engineers can achieve sufficient expression for high production while maintaining regulatory control to prevent toxicity.
Different production hosts may require distinct UTR design strategies based on their native genetic architecture. For example, bacterial hosts from the Actinobacteria group (which naturally contain >20% leaderless genes) [10] may express leaderless constructs more efficiently than traditional E. coli chassis. Understanding these host-specific differences enables better matching of genetic part to cellular context. Additionally, the development of hybrid, mutated, and artificial UTR libraries [67] provides a resource for optimizing expression across diverse industrial hosts and applications, from enzyme production to metabolic engineering of complex natural products.
Decision Framework and Future Perspectives
UTR Selection Algorithm
Selecting the appropriate UTR architecture for a specific synthetic biology application requires systematic consideration of multiple factors. The following decision framework outlines key considerations and recommended paths based on application requirements:
This decision framework emphasizes the importance of aligning UTR selection with specific application requirements. For maximum protein production in well-characterized hosts, strong leadered UTRs with optimized Shine-Dalgarno sequences and minimal secondary structure typically yield the highest expression levels [13] [67]. In contrast, applications requiring rapid response times or implementation in genetic circuits may benefit from leaderless architectures that eliminate the timing delays associated with ribosome scanning through 5' UTRs [10] [62]. For non-model organisms, where characterized UTR parts may be limited, library-based approaches that screen artificial or semi-artificial UTR variants provide a path to identifying functional sequences [67].
Emerging Technologies and Future Directions
The field of UTR engineering is being transformed by several emerging technologies that promise to enhance both the precision and efficiency of optimization. Machine learning approaches are increasingly being applied to predict UTR function from sequence, potentially reducing the experimental burden of library screening [67]. As these models improve, they may enable purely computational design of UTRs with specified regulatory properties. Additionally, massively parallel reporter assays continue to increase in scale and sophistication, providing comprehensive datasets that capture UTR function across multiple cellular conditions [68]. These resources will be invaluable for building predictive models that account for context-dependence.
Future advances will likely focus on expanding the toolkit of regulatory modalities available through UTR engineering. The discovery that "uORFs have been found to increase reinitiation with the longer distance between its uAUG and the start codon of the main ORF" [1] suggests opportunities for designing synthetic uORFs that provide precise translational control. Similarly, the development of small RNA-responsive UTRs could enable sophisticated genetic circuits that integrate multiple inputs. As synthetic biology applications continue to expand into diverse bacterial hosts, the need for host-specific UTR design principles will drive continued investigation into the fundamental mechanisms of translation initiation and regulation across the bacterial domain.
A Head-to-Head Comparison: Functional Outcomes in Gene Expression and Regulation
Translation initiation is the critical, rate-limiting step in protein synthesis, and its mechanisms are fundamentally divergent across the domains of life. Historically, textbook knowledge held a relatively simple dichotomy: in prokaryotes, the small ribosomal subunit (30S) binds directly to a Shine-Dalgarno (SD) sequence on leadered mRNAs, while in eukaryotes, the small subunit (40S) scans from the 5' cap to locate the start codon [69] [70]. However, contemporary research reveals a more complex and fascinating landscape, primarily driven by the study of leaderless genes. These genes, which lack any 5' untranslated region (5'-UTR), necessitate initiation mechanisms that do not rely on upstream signals, thereby challenging conventional models [2]. Research into leadered versus leaderless genes has been pivotal in uncovering novel initiation pathways, such as 70S-scanning in bacteria and internal ribosome entry in eukaryotes. This whitepaper provides an in-depth technical guide to these mechanisms, framing them within the broader thesis that leaderless genes represent an ancient and widespread initiation strategy whose study continues to refine our understanding of gene expression control. For researchers and drug development professionals, mastering these mechanisms is essential for applications ranging from the design of gene expression systems to the development of novel classes of antibiotics that target unique initiation pathways.
Core Mechanisms: A Comparative Analysis
Prokaryotic Initiation Mechanisms
In bacteria, three distinct initiation pathways have been characterized, each with specific factor requirements and functional roles.
The 30S-Binding Initiation (Canonical Leadered Initiation): This is the standard mechanism for leadered mRNAs. The 30S ribosomal subunit, with the help of three initiation factors (IF1, IF2, and IF3), binds to the mRNA. Recognition is mediated by base-pairing between the anti-Shine-Dalgarno (aSD) sequence at the 3' end of the 16S rRNA and the SD sequence upstream of the AUG start codon on the mRNA, positioning the ribosome correctly for initiation [69] [46]. The SD sequence's complementarity to the aSD and its spacing from the start codon are key determinants of translation initiation efficiency [46].
The 70S-Scanning Initiation (A Novel Leadered Mechanism): Recently demonstrated, this mechanism operates on polycistronic mRNAs. Evidence indicates that following translation of an upstream cistron, the 70S ribosome does not necessarily dissociate but rather scans the intercistronic region for the next SD sequence to initiate downstream translation [69]. This scanning is triggered by fMet-tRNA and does not require energy from GTP hydrolysis. Notably, this pathway has specific initiation factor requirements; IF3 is essential, and IF1 is highly stimulating, with the latter's role being to prevent untimely interference by elongator tRNA•EF-Tu•GTP complexes [69]. It is estimated that this novel mode accounts for at least 50% of bacterial initiation events, underscoring its significant biological role [69].
Leaderless Initiation (Non-Canonical): Leaderless mRNAs (lmRNAs) possess a 5' end that starts with or is very near the AUG initiation codon, thus lacking an SD sequence. These mRNAs can be directly bound by the intact 70S ribosome, a process that is conserved across bacteria, archaea, and eukaryotes [71] [2]. A striking feature of lmRNA initiation is its ability to occur, albeit inefficiently, in the absence of all three initiation factors [69]. Structural studies have shown that efficient translation of certain lmRNAs, such as the λcI repressor, is enhanced in ribosomal mutants deficient in protein uS2. The absence of uS2 leads to the loss of bS21, which normally supports the aSD region. This repositioning of the aSD case lmRNA exit from the ribosome, facilitating leaderless initiation [71].
Eukaryotic Initiation Mechanism
Eukaryotic translation initiation is predominantly governed by the scanning mechanism, a stark contrast to the direct binding observed in most prokaryotic pathways.
- The 40S Subunit Scanning Mechanism (Canonical): For the vast majority of eukaryotic mRNAs, initiation involves the attachment of the 40S ribosomal subunit, loaded with initiator tRNA and multiple initiation factors (collectively the 43S pre-initiation complex or PIC), to the 5' cap of the mRNA. This assembly is facilitated by the eIF4F complex and poly(A)-binding protein (PABP), potentially forming a "closed-loop" mRNA structure [70]. The PIC then scans the 5' UTR in a 5'-to-3' direction, inspecting each codon in a process that requires ATP and is aided by RNA helicases like eIF4A to unwind secondary structures [70]. The start codon is identified primarily by perfect complementarity between the AUG codon and the anticodon of the initiator tRNA. This recognition event triggers GTP hydrolysis, factor release, and joining of the 60S large subunit to form the elongation-competent 80S ribosome [70]. Recent transcriptome-wide studies in mouse brain tissue using techniques like RCP-seq have directly captured these scanning 40S subunits, showing their accumulation upstream of the start codon in a "poised" configuration, which correlates with enhanced translational efficiency [72].
Table 1: Comparative Analysis of Translation Initiation Mechanisms
| Feature | 30S-Binding (Prokaryotic) | 70S-Scanning (Prokaryotic) | Leaderless (Prokaryotic) | 40S Scanning (Eukaryotic) |
|---|---|---|---|---|
| Ribosome State | 30S subunit | 70S ribosome | 70S ribosome | 40S subunit (43S PIC) |
| mRNA Type | Leadered, polycistronic | Leadered, polycistronic | Leaderless (lmRNA) | Leadered, monocistronic |
| Key Recognition Signal | Shine-Dalgarno (SD) sequence | Shine-Dalgarno (SD) sequence | Start codon (AUG) itself | AUG codon in Kozak context |
| Initiation Factor Requirement | IF1, IF2, IF3 | IF3 (essential), IF1 (stimulatory) | Can be factor-independent | >12 factors (eIF2, eIF3, eIF4F, etc.) |
| Energy Requirement | GTP (via IF2) | Not required for scanning | Not well defined | ATP (helicases), GTP (eIF2) |
| Prevalence | Well-characterized, common | ~50% of events [69] | Widespread, up to 26% in some genomes [2] | Dominant mechanism |
| Evolutionary Context | Bacterial/Archaeal | Bacterial | Possibly ancestral (LUCA) [2] | Eukaryotic |
Experimental Methodologies and Visualization
Deciphering these complex initiation mechanisms relies on a suite of sophisticated biochemical, structural, and genomic techniques.
Key Experimental Protocols
In Vitro Reconstitution and Toe-Printing Assay: This classic biochemical approach was instrumental in characterizing the 70S-scanning mechanism [69]. The methodology involves:
- Reconstitution: Purified ribosomal subunits, initiation factors (IF1, IF2, IF3), fMet-tRNAfMet, and engineered monocistronic or bicistronic mRNA templates are incubated together in a physiological buffer to form initiation complexes.
- Reverse Transcription: An mRNA-specific DNA primer, radioactively or fluorescently labeled, is added to the complex.
- cDNA Synthesis: Reverse transcriptase extends the primer until it is sterically blocked ("toe-printed") by the stalled ribosome complex.
- Electrophoresis: The resulting cDNA fragments are resolved on a sequencing gel. The length of the truncated cDNA product precisely maps the leading edge of the ribosome, typically ~15-17 nucleotides downstream of the P-site codon, allowing researchers to determine the ribosome's position on the mRNA with nucleotide-level resolution [70].
Cryo-Electron Microscopy (Cryo-EM) for Structural Insight: Cryo-EM has provided atomic-level insights into the structure of initiation complexes, particularly for leaderless initiation. The protocol for studying the λcI lmRNA complex [71] is:
- Sample Preparation: Purified 70S ribosomes (from both wild-type and mutant rpsB11 E. coli strains deficient in uS2) are incubated with λcI lmRNA and fMet-tRNAfMet.
- Vitrification: The complex is rapidly frozen in thin ice on a cryo-EM grid.
- Data Collection: Millions of particle images are collected using an electron microscope.
- Computational Analysis: 2D classification, 3D refinement, and focused classification are used to resolve distinct structural populations (e.g., ribosomes with and without uS2/bS21).
- Model Building: Atomic models are built and refined to understand how the absence of specific ribosomal proteins alters the architecture of the mRNA exit channel, thereby facilitating lmRNA accommodation.
Ribosome Complex Profiling (RCP-seq): This nucleotide-resolution method maps the positions of small ribosomal subunits (SSUs) across the transcriptome to study scanning. The adapted protocol for mammalian brain tissue [72] involves:
- In Vivo Crosslinking: Mouse brain tissue (e.g., dentate gyrus) is irradiated with UV light to covalently crosslink ribosomes to their bound mRNAs.
- Cell Lysis and Nuclease Digestion: The tissue is lysed, and the lysate is treated with RNase I to digest unprotected RNA, leaving ribosome-protected mRNA fragments (footprints).
- Sucrose Gradient Centrifugation: The digest is fractionated via sucrose density gradient ultracentrifugation to separate 40S (SSU) and 80S (monosome) complexes.
- Library Preparation and Sequencing: RNA is extracted from the SSU and 80S fractions, and sequencing libraries are constructed.
- Bioinformatic Analysis: Sequencing reads are aligned to the transcriptome. The accumulation of SSU footprints immediately upstream of the start codon provides a direct snapshot of scanning PICs, allowing researchers to quantify "poised" ribosomes and study regulatory events during the initiation phase.
Visualization of Mechanisms and Workflows
Diagram 1: 70S scanning initiation in prokaryotes
Diagram 2: 40S subunit scanning in eukaryotes
Diagram 3: RCP-seq experimental workflow
The Scientist's Toolkit: Essential Research Reagents and Materials
Table 2: Key Research Reagent Solutions
| Reagent/Material | Function in Research | Specific Application Example |
|---|---|---|
| Purified Ribosomal Subunits & Factors | For in vitro reconstitution of translation initiation complexes. | Mechanistic studies of 70S-scanning by forming defined complexes with mutant factors [69]. |
| Engineered Bicistronic mRNA Templates | To study ribosomal behavior between coding sequences. | Demonstrating 70S-scanning and translational coupling [69]. |
| Antisense Oligo-DNA Blockers | To selectively inhibit translation or scanning of specific mRNA regions. | Blocking the first cistron or intercistronic region in bicistronic mRNA assays [69]. |
| Cryo-EM Equipment & Software | To determine high-resolution structures of ribosomal complexes. | Solving the structure of 70S ribosomes bound to leaderless mRNA [71]. |
| RCP-seq/TCP-seq Library Kits | For preparing sequencing libraries from ribosome-protected mRNA footprints. | Genome-wide mapping of scanning 40S subunits in complex tissues [72]. |
| uS2-Deficient Mutant Strains (e.g., rpsB11) | To study the role of specific ribosomal proteins in initiation. | Investigating enhanced leaderless mRNA translation in E. coli [71]. |
| Dual-Luciferase Reporter Assay Systems | For simultaneous, quantitative measurement of two cistrons' translation. | Quantifying the effect of scanning blockade on downstream cistron expression [69]. |
Discussion and Research Implications
The delineation of multiple translation initiation mechanisms, particularly through the lens of leaderless versus leadered genes, has profound implications for basic research and therapeutic development. The discovery that 70S-scanning initiation accounts for approximately half of all bacterial initiation events revolutionizes the traditional view of translation in prokaryotes and suggests a mechanism for efficient translational coupling in operons [69]. From an evolutionary standpoint, the conservation of leaderless initiation across all domains of life, and its prevalence in ancient bacterial phyla, provides compelling support for the hypothesis that it represents an ancestral mechanism employed by the last universal common ancestor (LUCA) [2]. The structural insights showing that minor alterations in ribosomal proteins (like uS2) can significantly modulate the efficiency of lmRNA translation reveal a layer of ribosomal specialization previously underappreciated [71].
For the field of drug development, these mechanistic differences represent a treasure trove of potential targets. The unique factor requirements of the 70S-scanning mode (e.g., essential IF3) and the distinct structure of lmRNA initiation complexes could be exploited to design next-generation antibiotics that selectively disrupt pathogenic bacterial translation without affecting the host eukaryotic machinery. Furthermore, the ability to map scanning ribosomes in tissues like the mammalian brain [72] opens new avenues for understanding and treating neurological disorders where dysregulated translation at synapses is a key factor. Continued research into these diverse initiation pathways will undoubtedly yield critical insights for both fundamental molecular biology and applied biomedicine.
In the complex cellular environment, organisms are continually subjected to various stressors that inhibit standard gene expression programs. The ability to maintain protein synthesis under these inhibitory conditions is a critical determinant of survival for cells across all domains of life. This adaptability is largely governed by the fundamental architecture of mRNA transcripts, which can be broadly categorized as either "leadered" or "leaderless." These structural differences dictate distinct translational mechanisms with profound implications for stress response and regulatory flexibility.
Leadered mRNAs, long considered the canonical standard, possess 5' untranslated regions (5' UTRs) containing regulatory elements such as the Shine-Dalgarno (SD) sequence in bacteria. In contrast, leaderless mRNAs (lmRNAs) completely lack 5' UTRs, with the start codon positioned at or extremely near the 5' end of the transcript [11]. Once regarded as molecular relics, lmRNAs are now recognized as significant components of transcriptomes across diverse taxa, comprising up to 25% of all transcripts in Mycobacterium tuberculosis and up to 60% in some bacterial species like Deinococcus deserti [11] [63]. This technical review examines the mechanistic differences in translation initiation between these mRNA classes and their specialized roles under stress conditions, providing experimental frameworks for investigating these pathways and their potential applications in therapeutic development.
Fundamental Mechanisms of Translation Initiation
Canonical Leadered Translation
The canonical translation initiation mechanism for leadered mRNAs employs well-defined sequential steps. In bacteria, the small ribosomal subunit (30S) binds the mRNA's 5' UTR through complementary base pairing between the anti-Shine-Dalgarno (aSD) sequence on 16S rRNA and the SD sequence upstream of the start codon [11]. This interaction is facilitated by initiation factors IF1, IF2, and IF3, with the RNA-binding protein bS1 assisting in unfolding structured 5' UTRs [11]. The 30S complex then recruits the large ribosomal subunit (50S) to form the functional 70S ribosome capable of elongation.
Eukaryotic leadered translation follows a distinct but analogous pathway where the 43S pre-initiation complex, comprising the small ribosomal subunit (40S) and multiple initiation factors, recognizes the 5' cap structure and scans the 5' UTR until it encounters the start codon [17]. Both mechanisms rely heavily on structured 5' UTR elements and specific initiation factors, making them vulnerable to disruptions in these components under stress conditions.
Alternative Initiation Pathways for Leaderless mRNAs
Leaderless mRNAs employ fundamentally different initiation mechanisms that bypass many requirements of canonical translation. Four distinct pathways have been identified for lmRNA translation in eukaryotes alone [17]:
- 80S ribosome scanning: Direct binding of assembled 80S ribosomes to the 5' terminal start codon without prior subunit dissociation [17]
- eIF2-dependent initiation: Canonical initiation factor-mediated assembly, though this is inefficient for lmRNAs [17]
- eIF2D-mediated mechanism: Utilization of alternative initiation factor eIF2D [17]
- eIF5B/IF2-assisted initiation: A bacterial-like pathway where the universal translation initiation factor eIF5B (bacterial IF2 ortholog) facilitates initiator tRNA recruitment [17]
In bacteria, lmRNA translation occurs primarily through direct binding of 70S ribosomes to the initiation codon, with IF2 playing a particularly important role in stabilizing the initiator tRNA and mRNA binding [11] [17]. This mechanism is notably independent of SD sequence interactions and shows reduced dependence on initiation factors under certain conditions [11].
Figure 1: Comparative translation initiation pathways for leadered and leaderless mRNAs. Leaderless initiation employs more direct mechanisms that contribute to enhanced stress resistance.
Differential Responses to Stress Conditions
Translation Under Nutrient Limitation and Oxidative Stress
The structural simplicity of leaderless mRNAs confers significant advantages under diverse stress conditions. Research in Mycobacterium tuberculosis demonstrates that leaderless translation remains robust during nutrient starvation and nitric oxide exposure, while leadered translation is significantly compromised [63]. In one study, luminescent reporter strains of M. tuberculosis containing either leadered (desA1) or leaderless (desA2) constructs showed markedly different responses to stress: leaderless translation was significantly more stable than leadered translation during adaptation to nutrient starvation and nitric oxide exposure [63]. Similar stability was observed during early macrophage infection, suggesting lmRNAs provide physiological advantages during host-pathogen interactions [63].
In eukaryotic systems, leaderless mRNA translation exhibits remarkable resistance to stressors that impair canonical initiation. The FLeeTING mRNA Transfection (FLERT) technique revealed that leaderless translation in mammalian cells remains efficient under arsenite-induced oxidative stress and dithiothreitol-induced unfolded protein stress, conditions that severely inhibit leadered translation [17]. This resistance stems from the reduced dependence of lmRNA translation on the eIF4F complex and eIF2, both primary targets of stress-induced translational control mechanisms [17].
Response to Translation-Specific Inhibitors
Chemical inhibition studies further highlight the mechanistic differences between these initiation pathways. Minocycline, a tetracycline antibiotic that attenuates cytoplasmic translation, extends lifespan and reduces protein aggregation even in post-stress-responsive C. elegans [73]. This effect occurs through preferential attenuation of highly translated mRNAs, disproportionately affecting leadered transcripts while preserving limited translation capacity that benefits lmRNAs.
Eukaryotic leaderless translation demonstrates partial resistance to harringtonine and T-2 toxin, elongation inhibitors that preferentially target de novo assembled 80S ribosomes [17]. At concentrations that completely inhibit leadered translation (0.1-0.2 μM), leaderless mRNA translation persists, suggesting different ribosomal conformations or factor requirements during initiation [17].
Table 1: Stress Response Characteristics of Leadered vs. Leaderless Translation
| Stress Condition | Leadered mRNA Response | Leaderless mRNA Response | Experimental System |
|---|---|---|---|
| Nutrient Starvation | Significant reduction in translation [63] | Maintained translation efficiency [63] | Mycobacterium tuberculosis reporter strains |
| Oxidative Stress (Arsenite) | Nearly complete inhibition at 20μM [17] | Partial resistance, continued translation [17] | Mammalian cells (FLERT assay) |
| Unfolded Protein Response (DTT) | Strong inhibition [17] | Pronounced resistance, 10x advantage [17] | Mammalian cells (FLERT assay) |
| mTOR Inhibition (Torin1) | Up to 4-fold inhibition [17] | Almost complete resistance [17] | Mammalian cells (FLERT assay) |
| Nitric Oxide Exposure | Significant reduction [63] | Sustained translation levels [63] | Mycobacterium tuberculosis reporter strains |
| Macrophage Infection | Transient reduction [63] | More stable translation [63] | Mycobacterium tuberculosis infection model |
Experimental Approaches and Methodologies
Reporter Systems for Monitoring Translation Efficiency
The construction and application of reporter systems is fundamental for quantifying differential translation activity. For investigating mycobacterial systems, the following protocol has been successfully employed [63]:
Plasmid Construction:
- Use integrating expression vectors (e.g., pMV306trpT) with strong transcriptional terminators to prevent read-through
- Clone firefly luciferase (ffluc) gene devoid of its optimized Shine-Dalgarno sequence
- Fuse candidate regulatory regions from representative genes to the 5' end of the reporter:
- For leadered constructs: Include 50bp promoter region + 5'UTR + six N-terminal amino acids from leadered gene (e.g., desA1/Rv0824c)
- For leaderless constructs: Include 50nt promoter region + six N-terminal amino acids from leaderless gene (e.g., desA2/Rv1094)
- Verify constructs by sequencing across cloning junctions
Bacterial Strain Generation and Analysis:
- Generate luminescent reporter strains by transforming constructs into target organisms (e.g., M. tuberculosis H37Rv)
- For translation measurements during stress:
- Grow cultures to exponential phase (OD~600~ ~0.5-1.0)
- Apply stress conditions: nutrient starvation in minimal media, nitric oxide donors (e.g., DETA/NO)
- Monitor luminescence at regular intervals alongside viability controls
- Normalize luminescence readings to bacterial density
- Confirm transcription levels via quantitative RT-PCR on parallel samples to distinguish translational from transcriptional effects
The FLeeTING mRNA Transfection (FLERT) Technique
For eukaryotic systems, the FLERT technique enables precise analysis of translation mechanisms in living cells while minimizing secondary effects [17]:
mRNA Preparation:
- Generate capped, polyadenylated reporter transcripts in vitro
- Experimental construct: Leaderless mRNA with 5'-terminal start codon (e.g., cI-derived sequence)
- Control constructs: Leadered mRNAs with structured 5'UTRs (e.g., β-actin UTR)
- Include internal control: Renilla luciferase (Rluc) mRNA with standard 5'UTR
Transfection and Stress Application:
- Culture mammalian cells in 24-well plates to 70-80% confluence
- Prepare mRNA mixtures containing experimental and control reporters
- Apply stress inducers immediately prior to transfection (5-minute pre-treatment):
- Oxidative stress: Sodium arsenite (20-500μM)
- Unfolded protein response: Dithiothreitol (1-5mM)
- mTOR inhibition: Torin1 (250nM)
- Transfect mRNA mixtures using appropriate transfection reagents
- Harvest cells 2 hours post-transfection for luciferase activity measurements
Data Analysis:
- Normalize firefly luciferase values to Renilla luciferase internal controls
- Calculate relative translation efficiency under stress vs. non-stress conditions
- Compare stress resistance between leaderless and leadered constructs
Table 2: Essential Research Reagents for Studying Translation Mechanisms
| Reagent/Category | Specific Examples | Function/Application | Key Characteristics |
|---|---|---|---|
| Reporter Vectors | pMV306trpT, pTC1 + P~desA1~:desA1', pTC1 + P~desA2~:desA2' [63] | Quantifying translation of different mRNA classes | Integrating vectors for stable maintenance; strong terminators to prevent read-through |
| Reporter Genes | Firefly luciferase (ffluc), Renilla luciferase (Rluc) [17] [63] | Sensitive measurement of translation efficiency | Rapid turnover for dynamic measurements; dual systems for normalization |
| Stress Inducers | Sodium arsenite, Dithiothreitol (DTT), Torin1, Nitric oxide donors [17] [63] | Imposing specific translational stresses | Well-characterized mechanisms; dose-dependent effects |
| Inhibitors | Harringtonine, T-2 toxin, Minocycline [17] [73] | Probing specific initiation mechanisms | Target distinct steps in translation; different sensitivity profiles |
| Cell Models | M. tuberculosis H37Rv, Cultured mammalian cells, C. elegans [17] [63] [73] | In vivo translation analysis | Representative of different biological systems; genetic tractability |
Evolutionary and Functional Significance
Distribution Across the Tree of Life
Leaderless genes represent an ancient translation initiation mechanism potentially predating the divergence of the three domains of life. Bioinformatic analysis of 953 bacterial and 72 archaeal genomes reveals that leaderless genes are widespread, though not dominant, across diverse bacterial taxa [10]. The proportion varies significantly between phylogenetic groups, with Actinobacteria and Deinococcus-Therpus containing over 20% leaderless genes in their genomes [10]. This distribution suggests an evolutionary trajectory where leadered translation has generally become more predominant, though lmRNAs remain functionally important in specific lineages.
The conservation of lmRNA translation mechanisms from bacteria to eukaryotes underscores their fundamental importance. Eukaryotes retain the capacity to translate lmRNAs through multiple pathways despite the predominance of scanning-based initiation [17]. This conservation suggests maintaining lmRNA translation capability provides selective advantages, particularly under stress conditions where canonical initiation is compromised.
Physiological Roles in Stress Adaptation
The persistence of leaderless translation systems appears linked to their specialized role in stress response networks. In M. tuberculosis, proteins with secondary adaptive functions, including toxin-antitoxin systems, are preferentially encoded by leaderless transcripts [63]. The ratio of leaderless to leadered transcripts increases during growth arrest, suggesting lmRNAs contribute to the non-replicating persistent state [63].
The non-autonomous regulation of stress responses through lmRNA translation adds another layer of biological significance. In C. elegans, germline-specific knockdown of cytochrome c (cyc-2.1) non-autonomously activates the intestinal mitochondrial unfolded protein response (UPR^mt^) and AMPK signaling, extending lifespan through a translationally regulated mechanism [74]. This demonstrates how tissue-specific lmRNA translation can coordinate organism-wide stress responses.
Figure 2: Stress response network showing how leaderless translation bypasses critical inhibition points to maintain proteostasis. While leadered translation is blocked by eIF2 phosphorylation and eIF4F inactivation, leaderless translation continues through alternative initiation mechanisms.
Implications for Therapeutic Development
The unique properties of leaderless mRNA translation present compelling opportunities for therapeutic intervention. The demonstrated resilience of lmRNA translation under stress conditions suggests strategic approaches for combating persistent bacterial infections. In Mycobacterium tuberculosis, where leaderless transcripts encode stress adaptation proteins, targeted inhibition of lmRNA translation could undermine bacterial persistence during antibiotic treatment [63].
In neurodegenerative disease, where protein aggregation and stress response deficiency converge, translation attenuation strategies inspired by lmRNA characteristics show promise. Minocycline extends lifespan and reduces protein aggregation even in post-stress-responsive C. elegans by preferentially attenuating highly translated mRNAs, effectively rebalancing the proteostasis network without activating stress signaling pathways [73]. This suggests that modulators of translational selectivity could bypass the age-related collapse of stress response activation.
The mechanistic insights from leaderless translation pathways also inform therapeutic mRNA design. The stress-resistant properties of lmRNAs could be leveraged to maintain therapeutic protein expression under pathological conditions where canonical translation is suppressed. This approach might be particularly valuable for cytoprotective gene therapies in neurodegenerative conditions, stroke, or ischemia-reperfusion injury where oxidative stress severely compromises cellular translation capacity.
The structural dichotomy between leadered and leaderless mRNAs represents a fundamental biological strategy for managing translation under diverse environmental conditions. While leadered translation provides sophisticated regulatory control during optimal growth, leaderless translation offers resilience when conventional initiation mechanisms are compromised. This division of labor enables organisms to maintain essential protein synthesis during stress through specialized mRNAs that bypass the most vulnerable steps of canonical initiation.
The experimental frameworks outlined here provide methodologies for quantifying these differential responses and investigating their mechanistic bases. As our understanding of these systems deepens, opportunities emerge for targeting these pathways therapeutically—either by disrupting bacterial persistence mechanisms or by maintaining protective gene expression in stressed eukaryotic cells. The continued investigation of these alternative translation mechanisms will undoubtedly yield further insights into cellular adaptation and new approaches for addressing complex diseases of protein homeostasis.
The stability of messenger RNA (mRNA) is a critical determinant of gene expression levels, influencing cellular adaptation, stress response, and pathogenesis. Unlike transcriptional control, which regulates mRNA synthesis, post-transcriptional regulation through mRNA turnover allows for rapid adjustment of protein output in response to changing environments. In bacterial systems, a fundamental distinction exists between leadered and leaderless gene architectures, which profoundly impacts their respective mRNA stability and translational efficiency.
Leadered transcripts contain 5' untranslated regions (5' UTRs) that harbor regulatory elements, including the Shine-Dalgarno (SD) sequence for ribosome binding. In contrast, leaderless mRNAs initiate directly at the start codon, lacking a 5' UTR entirely. This structural difference suggests potentially divergent mechanisms of degradation and stability control. Research in mycobacteria and other bacterial species reveals that leaderless transcripts are not rare anomalies but represent a significant portion of the transcriptome—approximately 14-25% of genes in Mycobacterium tuberculosis and Mycobacterium smegmatis [13] [14]. Understanding the differential stability characteristics between these transcript classes provides crucial insights into bacterial adaptation and opens novel avenues for therapeutic intervention.
Quantitative Comparison of mRNA Stability
Direct comparisons of half-life between leadered and leaderless transcripts reveal complex regulatory patterns influenced by multiple factors. Experimental data from model organisms provides quantitative insights into these relationships.
Table 1: Comparative mRNA Half-Life Characteristics
| Transcript Class | Organism | Key Features | Reported Half-Life | Influencing Factors |
|---|---|---|---|---|
| Leadered | Mycobacterium smegmatis (e.g., sigA 5' UTR) | Long 5' UTR (123 nt); contains SD sequence | Shorter half-life (instability conferred by long 5' UTR) [13] | 5' UTR secondary structure; RNase accessibility; transcription-translation coupling [13] |
| Leaderless | Mycobacterium smegmatis | Lacks 5' UTR; starts with start codon | Similar or comparable half-life to some leadered transcripts (e.g., those with sigA 5' UTR) [13] | Transcript production rate; absence of 5' end protection; ribosome binding efficiency [13] [14] |
| Aggregation-specific mRNAs | Dictyostelium discoideum | Developmentally regulated | >3 hours (in aggregated cells); 25-40 minutes (upon disaggregation) [75] | Cellular context; environmental cues; cell-cell contact [75] |
The stability of an mRNA is not an isolated property but is influenced by its own concentration. Studies in Escherichia coli and Lactococcus lactis demonstrate that increasing mRNA concentration can systematically reduce its stability, creating a negative feedback mechanism for gene regulation [76]. This inverse relationship appears to be a conserved physical mechanism across the bacterial kingdom, complicating direct comparisons between transcript classes.
Methodologies for Measuring mRNA Half-Life
Accurate determination of mRNA half-life is essential for understanding gene regulation. Several experimental approaches have been developed, each with specific applications and limitations.
Fluorescent Reporter Systems
Reporter systems using fluorescent proteins like yellow fluorescent protein (YFP) enable precise measurement of transcript stability under controlled genetic backgrounds [13].
- Principle: A promoter of interest drives the expression of a fluorescent protein fused to various 5' UTRs or leaderless sequences. By measuring fluorescence, protein abundance, mRNA abundance, and transcript half-life, researchers can calculate relative transcript production rates and infer stability.
- Workflow:
- Construct Design: Clone the 5' UTR or leaderless sequence of interest upstream of a reporter gene (e.g., yfp) in an expression plasmid.
- Transformation: Introduce the construct into the model organism (e.g., M. smegmatis).
- Culture & Sampling: Grow bacterial cultures and collect samples at multiple time points during exponential growth.
- Transcriptional Inhibition: Add a transcription inhibitor (e.g., rifampicin) to halt new RNA synthesis.
- mRNA Quantification: Isolve RNA from samples collected post-inhibition and quantify reporter mRNA levels using techniques like RT-qPCR or RNA-seq.
- Data Analysis: Plot remaining mRNA abundance over time and calculate the half-life from the decay curve.
Diagram 1: Reporter system workflow for measuring mRNA half-life.
RNA Metabolic Labeling
This approach involves direct chemical tagging of newly synthesized RNA to track its decay in real-time, providing high temporal resolution [77] [78].
- Principle: Incorporate labeled nucleotides (e.g., 4-thiouridine or 32PO4) into newly transcribed RNA. After a pulse of labeling, chase with unlabeled nucleotides and track the decay of the labeled RNA population over time.
- Workflow:
- Pulse Labeling: Expose cells to a medium containing the labeled nucleotide for a short, defined period.
- Chase Phase: Replace the labeling medium with an excess of unlabeled nucleotide medium.
- Time-Point Sampling: Collect cell aliquots at multiple time points after the chase begins.
- RNA Extraction & Purification: Isolve total RNA. For some labels (e.g., 4-thiouridine), biotinylate and purify the labeled RNA from the total RNA pool.
- High-Throughput Sequencing: Prepare sequencing libraries from the purified RNA samples.
- Bioinformatic Analysis: Map sequencing reads to the genome, quantify transcript abundance at each time point, and model decay kinetics to compute half-lives for each transcript.
Diagram 2: Metabolic labeling workflow for genome-wide half-life studies.
Mechanisms Influencing mRNA Stability
The degradation rate of an mRNA is governed by a complex interplay of cis-acting elements within the transcript itself and trans-acting factors within the cell.
1Cis-Acting Elements: The Role of the 5' End
The architecture of the 5' terminus is a primary determinant of mRNA stability.
- Leadered Transcripts: The 5' UTR can significantly modulate stability. For instance, the long 5' UTR of M. smegmatis sigA confers instability, leading to a shorter half-life. This can be attributed to:
- Secondary Structures: Stem-loop structures can protect the 5' end from exonucleases or hide internal cleavage sites from endonucleases [13] [76].
- Specific Sequences: Binding sites for ribonucleases or regulatory proteins (e.g., CsrA) within the UTR can promote or inhibit decay [13] [76].
- Translation Efficiency: Impairments in translation, often caused by strong secondary structures obscuring the RBS, can lead to faster mRNA decay due to reduced ribosome protection [13] [76].
- Leaderless Transcripts: Lacking a 5' UTR, these transcripts are missing a layer of regulatory information. Their stability is thus governed more directly by the coding sequence itself and the initiation of translation. In mycobacteria, leaderless transcripts are translated robustly and can exhibit stability comparable to leadered transcripts, though they often have lower predicted transcript production rates [13] [14].
2Trans-Acting Factors: The Degradation Machinery
A suite of ribonucleases and regulatory proteins executes mRNA decay.
- Major Ribonucleases:
- RNase E (in E. coli and mycobacteria): A key endonuclease that often initiates decay by internal cleavage, particularly preferring monophosphorylated 5' ends generated by the pyrophosphohydrolase RppH [76].
- RNase J (in B. subtilis and others): A bifunctional enzyme with both endonuclease and 5'-exonuclease activities [76].
- RNase Y: Essential in some Gram-positive bacteria for initiating degradation [76].
- The Exosome Complex: A multi-protein complex with exo- and endonuclease activity present in both eukaryotes and bacteria (e.g., Rrp6 in yeast) that degrades transcripts in the nucleus and cytoplasm [77].
- Regulatory RNA-Binding Proteins:
- Hfq: Facilitates the interaction between small regulatory RNAs (sRNAs) and their target mRNAs, often leading to degradation or translational repression [76].
- CsrA: A global regulator that binds to the 5' UTR of numerous mRNAs, affecting their stability, translation, and/or transcription elongation [76].
Diagram 3: Factors influencing mRNA stability.
The Scientist's Toolkit: Essential Research Reagents
The following table details key reagents and tools used in the featured experiments for studying mRNA stability.
Table 2: Essential Research Reagents for mRNA Stability Studies
| Reagent/Tool | Function | Example Use Case |
|---|---|---|
| Fluorescent Reporter Plasmids | Plasmid vectors containing promoter-UTR sequences fused to fluorescent protein genes (e.g., YFP). | Quantifying the effect of specific 5' UTRs or leaderless architectures on mRNA half-life and translation efficiency in vivo [13]. |
| Transcriptional Inhibitors | Small molecules that block RNA polymerase. | Halting new transcription to isolate and measure the decay of pre-existing mRNA (e.g., rifampicin) [13]. |
| Metabolic RNA Labels | Modified nucleotides (e.g., 4-thiouridine, 32PO4) incorporated into newly synthesized RNA. | Pulse-chase experiments to track the fate and decay kinetics of a defined cohort of mRNAs [75] [78]. |
| RNA Degradosome Mutants | Strains with deletions or mutations in genes encoding ribonucleases (e.g., rrp6∆, rnase E-) or associated factors. | Elucidating the role of specific degradation pathways in mRNA turnover and stability control [76] [77]. |
| Computational Tools (e.g., RNAtracker) | Software for analyzing RNA sequencing data from metabolic labeling experiments. | Distinguishing whether changes in mRNA abundance are due to altered transcription or decay rates, and identifying genetic variants affecting stability [78]. |
Implications for Drug Development and Therapeutics
Understanding mRNA stability, particularly the distinctions between leadered and leaderless transcripts, has profound implications for therapeutic development.
- Targeting Bacterial Adaptation: In pathogens like M. tuberculosis, which relies heavily on post-transcriptional regulation to adapt to host stress, the unique mechanism of leaderless translation presents a potential therapeutic target. Disrupting the stability or translation of key leaderless virulence factors could undermine bacterial survival [13] [14].
- Design of mRNA Therapeutics: The principles of mRNA stability are directly applied in the design of mRNA vaccines and therapies. Features such as 5' cap analogs, optimized 5' UTRs, and nucleoside modifications (e.g., pseudouridine, N1-methylpseudouridine) are incorporated into synthetic mRNAs to enhance their stability and translational capacity while reducing immunogenicity [60].
- Interpreting Disease Risk: Genetic variants that alter mRNA stability can influence disease susceptibility. Tools like RNAtracker help identify such variants, revealing that instability in mRNAs related to the innate immune system is linked to autoimmune diseases like lupus and multiple sclerosis [78]. This expands the potential targets for novel drugs aimed at modulating mRNA stability.
The analysis of global protein-to-mRNA ratios represents a critical frontier in systems biology, seeking to decipher the complex relationship between transcript abundance and the resulting protein synthesis that ultimately executes cellular functions. This relationship is surprisingly variable across genes and organisms, with mRNA levels often failing to perfectly predict protein abundance due to multi-layered post-transcriptional regulation [79]. This technical guide examines these relationships within a crucial contextual framework: the fundamental distinction between leadered and leaderless gene architectures.
In prokaryotes, a significant number of genes lack a 5' untranslated region (5' UTR); these are termed "leaderless" transcripts. Unlike traditional "leadered" transcripts that utilize a Shine-Dalgarno (SD) sequence within the 5' UTR for ribosome binding, leaderless transcripts initiate translation directly at the 5' end start codon [12]. Research indicates that approximately 14% of genes in mycobacteria such as Mycobacterium tuberculosis and Mycobacterium smegmatis are leaderless, a prevalence notably higher than in model organisms like E. coli [13] [10]. This architectural difference is not merely structural but has profound implications for how gene expression is regulated at the levels of transcription, translation, and mRNA decay, thereby directly influencing the protein-to-mRNA ratio [13]. Understanding these distinct regulatory paradigms is essential for accurately interpreting omics data and engineering microbial strains for synthetic biology and drug development.
Core Concepts: Leadered vs. Leaderless Genes
Fundamental Structural and Mechanistic Differences
The initiation of protein synthesis differs fundamentally between leadered and leaderless genes, setting the stage for divergent regulatory outcomes.
Leadered Genes represent the canonical initiation mechanism in bacteria. Their transcripts possess a 5' Untranslated Region (5' UTR) upstream of the start codon. This UTR typically contains a Shine-Dalgarno (SD) sequence that base-pairs with the anti-SD sequence on the 16S rRNA of the 30S ribosomal subunit, facilitating proper positioning of the ribosome at the start codon [12]. Assembly of the 70S ribosome occurs at the SD sequence, followed by translation initiation at the downstream AUG codon, typically with an N-terminal formylated methionine [12]. Experimentally, leadered genes produce nested RNA-seq and ribosome profiling (Ribo-seq) reads, where Ribo-seq reads begin downstream of the RNA-seq reads, reflecting the presence of the untranslated leader sequence [12].
Leaderless Genes represent a distinct and prevalent class in many bacteria. They lack a 5' UTR entirely, with the transcription start site (TSS) being identical to the translation initiation site. This structure means there is no Shine-Dalgarno sequence to guide the 30S subunit [12]. Instead, assembled 70S ribosomes are thought to bind directly to the 5' end of the mRNA and engage the start codon [13] [12]. This initiation mechanism is considered ancient, potentially used by the last universal common ancestor (LUCA), and is conserved across all domains of life [10]. In omics data, leaderless genes are identified by coincident 5' boundaries for RNA-seq and Ribo-seq reads, with the 5' triplet almost always being an AUG or GUG start codon [12].
Table 1: Comparative Features of Leadered and Leaderless Genes
| Feature | Leadered Genes | Leaderless Genes |
|---|---|---|
| 5' UTR | Present (median ~48-56 nt in mycobacteria) | Absent [13] [12] |
| Shine-Dalgarno Sequence | Typically present | Absent [12] |
| Ribosome Initiation Complex | 30S subunit binding, then 70S assembly | Pre-assembled 70S ribosome [12] |
| Transcription/Translation Start | Separate | Coincident [12] |
| Experimental Signature (Ribo-seq/RNA-seq) | Nested 5' boundaries | Coincident 5' boundaries [12] |
| Prevalence in Mycobacteria | ~86% of genes | ~14% of genes [13] |
The Protein-to-mRNA Ratio as a Functional Metric
The Protein-to-mRNA Ratio (ptr) is a quantitative measure that reflects the combined efficiency of all post-transcriptional processes for a given gene, culminating in protein synthesis. It is a key descriptor in systems biology models. A highly conserved ptr for a gene across different conditions or even species suggests that its expression is under tight, optimized control, often observed for essential cellular functions [80]. Conversely, a variable ptr indicates a gene subject to dynamic regulatory influences.
The relationship between mRNA and protein abundance is positive but not perfectly correlated. Studies across diverse bacteria and archaea show that mRNA levels typically explain only about 27% (with a range of 18-38%) of the variability in protein levels [80]. This discrepancy arises because protein abundance is influenced by a multitude of factors independent of mRNA concentration, including:
- Translation Efficiency: Governed by initiation rates, elongation rates, and codon usage [81] [82].
- mRNA Stability: The half-life of the transcript, which can be influenced by the 5' UTR structure, coding sequence, and RNase activity [13] [82].
- Protein Degradation: The relative stability of the synthesized protein [79].
The ptr is not merely a descriptive statistic; it has practical utility. Recent research demonstrates that RNA-to-protein (RTP) conversion factors can be derived from conserved ptr values. These factors allow for significantly improved prediction of protein abundance from transcriptomic data alone, a powerful tool for interpreting gene expression in complex microbial communities or experimental settings where proteomic measurement is challenging [80].
Key Analytical and Experimental Methodologies
Global Quantification of mRNA and Protein
A robust analysis of protein-to-mRNA ratios depends on high-quality, simultaneous measurements of the transcriptome and proteome.
Transcriptomics (e.g., RNA-seq)
- Purpose: To globally quantify mRNA abundance and identify transcription start sites (TSSs), which is critical for classifying genes as leadered or leaderless.
- Key Consideration: Specialized libraries, such as those capturing the 5' ends of transcripts, are required for precise TSS mapping, which definitively identifies leaderless transcripts (TSS = start codon) [13] [12].
Proteomics (Mass Spectrometry-based)
- Purpose: To globally identify and quantify protein abundances.
- Workflow: Proteins are digested into peptides, which are separated by liquid chromatography and analyzed by tandem mass spectrometry (MS/MS). Quantification is achieved through label-free methods or using isotopic labels (e.g., SILAC, TMT) [80].
Ribosome Profiling (Ribo-seq)
- Purpose: To provide a snapshot of the locations of actively translating ribosomes at nucleotide resolution.
- Protocol: Cells are treated with a translation inhibitor to stall ribosomes, followed by nuclease digestion of unprotected mRNA. The protected mRNA fragments ("ribosome footprints") are purified and sequenced [12].
- Utility for Architecture Studies: Ribo-seq uniquely distinguishes leadered and leaderless translation. Leaderless genes show ribosome footprints beginning precisely at the start codon, while leadered genes show footprints offset by the length of the 5' UTR [12].
The following diagram illustrates the core experimental workflow for generating and integrating multi-omics data to study protein-to-mRNA relationships.
Reporter Assays for Dissecting Expression Dynamics
While global omics methods identify correlations, reporter assays enable direct, causal testing of how specific genetic elements, like a 5' UTR, regulate expression. A common approach is to fuse the regulatory element of interest (e.g., the native sigA 5' UTR or a leaderless start) to a fluorescent reporter gene (e.g., YFP) expressed from a constitutive promoter [13].
This methodology allows for the independent quantification of the three key facets of gene expression that collectively determine the protein-to-mRNA ratio:
- Transcript Production Rate: Measured using qPCR or RNA-seq on the reporter transcript and calculated from steady-state mRNA abundance and half-life.
- mRNA Half-Life: Determined by treating cells with a transcription inhibitor (e.g., rifampin) and tracking the decay of the reporter mRNA over time using qPCR [13].
- Translation Efficiency and Protein Abundance: Quantified directly by measuring fluorescence and protein yield.
Table 2: Key Experimental Findings from Leadered and Leaderless Reporter Studies
| Experimental Manipulation | Impact on Transcript Production Rate | Impact on mRNA Half-Life | Impact on Translation Efficiency |
|---|---|---|---|
| Long 5' UTR (e.g., sigA) | Increased [13] | Decreased (shorter half-life) [13] | Decreased [13] |
| Leaderless Architecture | Lower than leadered [13] | Similar to sigA 5' UTR [13] | Similar to or lower than leadered (context-dependent) [13] |
| Control Synthetic 5' UTR | Baseline | Baseline (longer half-life) [13] | Baseline (higher) [13] |
Research Reagent and Experimental Solutions
Successful execution of these experiments relies on a suite of specialized reagents and tools. The following table details key components for a researcher's toolkit.
Table 3: Research Reagent Solutions for Protein-to-mRNA Studies
| Reagent / Tool | Function / Utility | Example Application |
|---|---|---|
| Fluorescent Protein Reporters (e.g., YFP, eGFP) | Enable quantitative, high-throughput measurement of protein expression levels in live cells. | Reporter constructs for testing 5' UTR function and translation efficiency [13]. |
| Nucleotide Analogues & Inhibitors | Arrest transcription or translation to measure kinetic parameters like mRNA half-life. | Rifampin (transcription inhibitor) for mRNA decay assays [13]. |
| Specialized Spacers (e.g., Fluor-PEG Puro) | Improve efficiency of mRNA-protein fusion for techniques like mRNA display. | Single-strand ligation for creating stable mRNA templates for in vitro selection [83]. |
| Mycobacterial Model Systems (e.g., M. smegmatis) | Non-pathogenic surrogate for studying gene regulation in a biologically relevant context. | Model organism for investigating leaderless translation and stress response in mycobacteria [13]. |
| Defined Genetic Elements (e.g., 5' UTR libraries) | Allow for systematic dissection of cis-regulatory sequences. | Testing the impact of the sigA 5' UTR on expression dynamics [13]. |
| Conserved RTP Conversion Factors | Gene-specific factors derived from conserved ptr ratios to predict protein from mRNA data. | Cross-species and cross-domain prediction of protein abundance from transcriptomic data [80]. |
Data Interpretation and Computational Integration
Elongation Dynamics and Codon Optimality
Regulation of protein abundance extends beyond initiation. The early elongation phase, particularly the identity of codons 3 to 5, significantly impacts protein yield. This effect is independent of tRNA abundance, translation initiation efficiency, or overall mRNA structure [81]. Single-molecule measurements reveal that ribosomes can pause or abort translation on these early codons, and introducing preferred sequence motifs in this region can enhance recombinant protein synthesis efficiency [81]. Furthermore, the ribosome itself controls mRNA stability in a codon-dependent manner, a phenomenon termed codon optimality. Codons decoded by abundant tRNAs (optimal codons) generally promote efficient elongation and mRNA stability, while rare codons can lead to ribosome pausing and mRNA decay [82].
A Cross-Domain Framework for Protein Prediction
A significant advancement in the field is the discovery that protein-to-mRNA (ptr) ratios for many orthologous genes are conserved across diverse bacteria and even between bacteria and archaea [80]. This conservation enables the calculation of RNA-to-protein (RTP) conversion factors from one well-studied organism to predict protein abundance in another, even distantly related, organism using only mRNA-seq data. This framework dramatically improves functional inference in complex microbiomes where proteomic data is unavailable [80]. The following diagram visualizes this cross-domain prediction concept.
The analysis of global protein-to-mRNA ratios reveals the intricate and multi-layered regulation of gene expression. Framing this analysis within the context of leadered versus leaderless gene architectures provides a powerful, mechanistic understanding of why these ratios vary. Leaderless genes, with their distinct initiation mechanism and regulatory constraints, represent a significant and under-explored paradigm in prokaryotic biology, especially in pathogens like M. tuberculosis.
Future research will focus on elucidating the precise molecular mechanisms that define the ptr of individual genes, particularly how the nascent peptide sequence during early elongation communicates with the ribosome exit tunnel to influence efficiency. Furthermore, the expansion and refinement of cross-domain RTP conversion factor libraries will unlock deeper insights from the vast quantities of existing and future transcriptomic data, bridging the gap between gene expression measurement and functional protein output. This knowledge is paramount for advancing fundamental microbiology, developing novel antibacterial strategies, and optimizing microbial systems for industrial and therapeutic applications.
The study of bacterial gene expression has been profoundly shaped by research in model organisms like Escherichia coli. However, over-reliance on such models can obscure unique biological mechanisms present in other bacteria. A 2025 analysis revealed that nearly 74% of bacterial species have never been the subject of a publication, while 50% of all articles focus on just 10 species, with E. coli dominating the field [84]. This taxonomic bias highlights the critical need to study non-model organisms to fully appreciate the diversity of microbial life.
Mycobacterium tuberculosis, the causative agent of tuberculosis, presents a compelling case study. This pathogen kills 1.5 million people globally each year and exhibits unique gene regulation mechanisms that enable its survival within human hosts [13] [27]. Unlike E. coli, mycobacteria express approximately 14% of their genes as leaderless transcripts [13], which completely lack 5' untranslated regions (5' UTRs). This fundamental difference in gene structure has profound implications for how these pathogens regulate gene expression in response to stress.
This review contrasts the mechanisms of gene regulation in mycobacteria, particularly focusing on leadered versus leaderless genes, with those of established model organisms like E. coli. We provide a comprehensive analysis of the distinctive features of mycobacterial gene expression, experimental approaches for their study, and the implications for drug development against mycobacterial diseases.
Fundamental Differences in Gene Structure and Regulation
Prevalence and Processing of Leaderless mRNAs
Leaderless mRNAs represent a fundamental divergence in genetic architecture between mycobacteria and traditional model organisms. These transcripts initiate directly at the start codon, lacking the 5' untranslated regions that contain ribosome-binding sites in conventional genes.
Table 1: Comparison of Leaderless mRNA Features in Bacteria
| Feature | Mycobacterium | Escherichia coli |
|---|---|---|
| Percentage of Transcriptome | ~14% of genes [13] | Rare under normal conditions [85] |
| Primary Function | Normal gene expression [13] | Stress response [85] |
| Translation Machinery | Standard 70S ribosomes [13] | Specialized "stress-ribosomes" [85] |
| Start Codon Recognition | Direct recognition by ribosomes [12] | Requires 5' AUG and modified 16S rRNA [85] |
| Translation Efficiency | Similar to leadered genes [13] | Less efficient than leadered genes [13] |
In E. coli, leaderless mRNAs typically emerge during stress conditions through the action of toxin-antitoxin systems. For example, the MazF toxin is an endoribonuclease induced under stress that cleaves single-stranded mRNAs at ACA sequences. When cleavage occurs at or near the start codon, leaderless mRNAs are generated [85]. Simultaneously, MazF also processes 16S rRNA, removing 43 nucleotides from the 3' terminus, including the anti-Shine-Dalgarno sequence. This generates specialized "stress-ribosomes" that preferentially translate the newly formed leaderless mRNAs [85].
In contrast, mycobacteria naturally maintain a high proportion of leaderless transcripts in their genome without requiring stress-induced modification of the translation machinery [13]. This suggests that leaderless translation is an integral, programmed component of normal gene expression in mycobacteria rather than primarily a stress response mechanism as in E. coli.
Translation Initiation Mechanisms
The molecular mechanisms for translation initiation differ significantly between leadered and leaderless mRNAs, and these differences have particular implications in mycobacteria.
Diagram 1: Translation initiation pathways for leadered and leaderless mRNAs
For leadered mRNAs, the 5' UTR contains a Shine-Dalgarno (SD) sequence that base-pairs with the anti-Shine-Dalgarno (aSD) sequence at the 3' end of 16S rRNA. This interaction facilitates the binding of the 30S ribosomal subunit upstream of the start codon, with initiation factor IF3 playing a crucial role in initiation complex formation [86]. The ribosome then scans to locate the start codon.
For leaderless mRNAs, a distinct mechanism applies. Research in E. coli has demonstrated that leaderless mRNAs bind directly to 70S ribosomes (rather than 30S subunits) in a process that requires initiator tRNA but is independent of IF3 [86]. The 5'-terminal AUG codon itself is both necessary and sufficient for ribosome binding, as demonstrated by experiments showing that adding a 5' AUG to a random RNA fragment renders it competent for ribosome binding [86]. Cross-linking studies using 4-thiouridine substituted at the +2 position of the AUG start codon revealed that leaderless mRNA forms tRNA-independent contacts with a subset of 30S subunit ribosomal proteins, suggesting initial interactions occur before tRNA stabilization [86].
In mycobacteria, leaderless transcripts appear to be translated robustly, in contrast to E. coli where they are generally less efficient [13]. Global comparisons in M. tuberculosis have failed to reveal systematic differences in protein/mRNA ratios for leadered versus leaderless transcripts, suggesting that translation efficiency variability is largely driven by factors other than leader status in mycobacteria [13] [27].
Experimental Approaches and Model Systems
Mycobacterium smegmatis as a Model Organism
Mycobacterium smegmatis, particularly the strain mc²155, has emerged as the primary model organism for mycobacterial research due to its non-pathogenicity, rapid growth (colonies in 3 days versus 3-4 weeks for M. tuberculosis), and high transformability [87]. This strain was isolated in 1990 as a mutant capable of efficient plasmid transformation, revolutionizing mycobacterial genetics [87].
Comparative genomics validates M. smegmatis as an excellent model for mycobacterial research. Of the ~4,000 protein-coding genes in M. tuberculosis, >2,800 have orthologs in M. smegmatis with >50% amino acid identity [87]. Essential gene sets are also well conserved - 96% of genes essential in M. smegmatis have orthologs in M. tuberculosis, and 90% of these are also essential in the pathogen [87]. This high conservation extends to core biological processes, including the unusual prevalence of leaderless transcripts.
Table 2: Model Organisms for Bacterial Gene Regulation Studies
| Characteristic | Mycobacterium smegmatis | Escherichia coli |
|---|---|---|
| Growth Rate | Fast (3 days for colonies) [87] | Very fast (overnight) [84] |
| Pathogenicity | Non-pathogenic [87] | Generally non-pathogenic (lab strains) [84] |
| Genetic Tools | Well-developed [87] | Extensive [84] |
| Relevance to Pathogens | High conservation with mycobacterial pathogens [87] | Limited for mycobacterial pathogens |
| Leaderless mRNA Prevalence | ~14% of transcriptome [13] | Minimal under normal conditions [85] |
| Drug Target Conservation | High with M. tuberculosis [87] | Low for anti-TB drugs |
Key Methodologies for Studying Leadered and Leaderless Genes
Reporter Constructs for Measuring Gene Expression Parameters
Critical insights into leadered and leaderless gene expression have come from carefully designed fluorescence reporter systems in M. smegmatis. The core methodology involves:
Construct Design: Creating fluorescent protein (e.g., YFP) reporters under the control of different 5' regulatory regions [13]. These include:
- Synthetic 5' UTRs (control)
- Native 5' UTRs from genes of interest (e.g., sigA)
- Leaderless configurations (no 5' UTR)
Parameter Measurement:
- Protein abundance: Measured via fluorescence intensity or immunodetection
- mRNA abundance: Quantified using RNA extraction and qRT-PCR
- mRNA half-life: Determined through transcriptional inhibition time courses
- Transcript production rates: Calculated from mRNA abundance and half-life data [13]
A key application of this approach involved investigating the sigA 5' UTR, which is unusually long (123 nt in M. smegmatis) and associated with a relatively short-lived mRNA [13]. Reporter constructs revealed that the sigA 5' UTR confers an increased transcript production rate, shorter mRNA half-life, and decreased apparent translation rate compared to a synthetic control 5' UTR [13] [27].
Ribosome Binding Assays
Toeprinting (primer extension inhibition) assays provide crucial information about ribosome-mRNA interactions:
- Complex Formation: Incubate ribosomes (70S or 30S subunits) with leaderless mRNA and initiator tRNA
- Primer Extension: Add reverse transcriptase and labeled primer; ribosomes block elongation
- Fragment Analysis: Resolve cDNA fragments via gel electrophoresis to map ribosome positions [86]
These assays demonstrated that leaderless mRNA binding to E. coli ribosomes is tRNA-dependent and requires a 5'-terminal AUG for stable binding [86]. The presence of a 5' AUG triplet alone can render random RNA fragments competent for ribosome binding, highlighting the importance of the start codon in leaderless translation initiation.
Cross-Linking Studies
Molecular interactions between leaderless mRNAs and ribosomes have been characterized using cross-linking approaches:
Probe Incorporation: A 4-thiouridine (4S-U) residue is incorporated at the +2 position of the AUG start codon in a model leaderless mRNA (e.g., from bacteriophage λ cI gene) [86]
Complex Formation and UV Activation: The modified mRNA is bound to ribosomes and cross-linked via UV irradiation
Interaction Mapping: Cross-linked rRNA and ribosomal proteins are identified through biochemical and mass spectrometry techniques [86]
These studies revealed that leaderless mRNA forms tRNA-independent contacts with specific 30S subunit ribosomal proteins, suggesting initial binding occurs before tRNA stabilization [86].
Table 3: Essential Research Reagents for Mycobacterial Gene Regulation Studies
| Reagent/Resource | Function/Application | Examples/Sources |
|---|---|---|
| M. smegmatis mc²155 strain | Non-pathogenic, fast-growing model for mycobacterial research | From laboratory stock collections [87] |
| Fluorescent reporter plasmids | Measure protein abundance, translation efficiency | Custom constructs with YFP/mCherry [13] |
| Shuttle phasmids | Genetic tools that replicate as plasmids in E. coli and phages in mycobacteria | TM4- and L1-based vectors [87] |
| Episomal plasmids | Gene expression, mutant complementation | pAL5000-based vectors [87] |
| Bioinformatic resources | Genomic analysis, ortholog identification | Mycobrowser, BioCyc [87] |
| 4-thiouridine (4S-U) | Photoactivatable nucleotide for RNA-protein cross-linking studies | Commercial suppliers [86] |
| Specialized ribosomes | Study translation initiation mechanisms | MazF-modified stress-ribosomes [85] |
Implications for Drug Development
The unique features of mycobacterial gene regulation present both challenges and opportunities for therapeutic development. Two first-line tuberculosis drugs, isoniazid and ethambutol, are active against M. smegmatis but not E. coli, enabling identification of their physiological targets using this model system [87]. Furthermore, Bedaquiline, the first new TB drug in 40 years, was discovered through a screening approach using M. smegmatis [87].
The prevalence of leaderless transcripts in mycobacteria suggests potential novel drug targets. Unlike E. coli where leaderless translation is primarily a stress response, in mycobacteria it represents a core component of gene expression. Species-specific differences in translation initiation mechanisms might be exploited to develop antibiotics that selectively disrupt mycobacterial protein synthesis without affecting host cells or beneficial microbiota.
M. smegmatis continues to serve as the vanguard for mycobacterial research, providing insights that would be difficult or impossible to obtain working directly with slow-growing pathogens. With the establishment of centralized resources like the Mycobacterial Systems Resource, this model organism will continue to accelerate discovery in the field [87].
The contrast between mycobacteria and traditional model organisms like E. coli reveals fundamental differences in genetic architecture, particularly in the prevalence and regulation of leaderless genes. Where E. coli largely employs leaderless transcripts as a specialized stress response, mycobacteria have integrated them as core components of their gene expression repertoire. These taxonomic differences underscore the importance of studying diverse bacterial systems rather than relying exclusively on traditional models.
Research in model mycobacteria like M. smegmatis has been instrumental in elucidating these mechanisms, providing genetic tractability while maintaining biological relevance to important pathogens. The continued development of tools and resources for mycobacterial research will undoubtedly yield further insights into the unusual gene regulation strategies of these important bacteria, potentially revealing novel targets for therapeutic intervention against tuberculosis and other mycobacterial diseases.
Conclusion
The dichotomy between leadered and leaderless genes represents a fundamental layer of complexity in gene regulation, with profound implications for understanding bacterial adaptation and virulence. The existence of multiple, parallel translation initiation pathways for leaderless mRNAs underscores a remarkable evolutionary flexibility. For biomedical research, the distinct regulatory patterns of leaderless genes, particularly their prevalence in pathogens like Mycobacterium tuberculosis, open promising avenues for therapeutic intervention. Future work should focus on elucidating the precise molecular triggers that favor one initiation mechanism over another and exploiting the unique features of leaderless transcription for developing novel antibiotics that disrupt a pathogen's adaptive response without affecting host cells. The integration of sophisticated computational predictions with robust experimental validation will be crucial to fully unravel the biological and clinical significance of these ancient genetic structures.
References
- 1. Five prime untranslated region [https://en.wikipedia.org/wiki/Five_prime_untranslated_region]
- 2. Leaderless genes in bacteria: clue to the evolution of ... [https://bmcgenomics.biomedcentral.com/articles/10.1186/1471-2164-12-361]
- 3. Differences in 5'untranslated regions highlight the ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11057154/]
- 4. Functional 5′ UTR mRNA structures in eukaryotic translation ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC5820134/]
- 5. Differences in 5'untranslated regions highlight the importance ... [https://genomebiology.biomedcentral.com/articles/10.1186/s13059-024-03248-0]
- 6. Human 5′ UTR design and variant effect prediction from a ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC7100133/]
- 7. High-throughput 5′ UTR engineering for enhanced ... [https://www.nature.com/articles/s41467-021-24436-7]
- 8. Comprehensive quantitative modeling of translation ... [https://link.springer.com/article/10.15252/msb.202211301]
- 9. Untranslated Regions (5′UTRs) in Zymomonas mobilis as ... [https://www.frontiersin.org/journals/microbiology/articles/10.3389/fmicb.2017.02432/full]
- 10. Leaderless genes in bacteria: clue to the evolution of ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC3160421/]
- 11. Regulation of Leaderless mRNA Translation in Bacteria [https://pmc.ncbi.nlm.nih.gov/articles/PMC9031893/]
- 12. Leaderless Transcripts and Small Proteins Are Common ... [https://journals.plos.org/plosgenetics/article/figures?id=10.1371/journal.pgen.1005641]
- 13. The Impact of Leadered and Leaderless Gene Structures ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC7148126/]
- 14. Leaderless Transcripts and Small Proteins Are Common ... [https://journals.plos.org/plosgenetics/article?id=10.1371/journal.pgen.1005641]
- 15. Molecular 'Fossils' Offer Microscopic Clues to the Origins ... [https://news.gatech.edu/news/2025/09/16/molecular-fossils-offer-microscopic-clues-origins-life-they-take-care-interpret]
- 16. Differential inhibition of 30S and 70S translation initiation ... [https://www.sciencedirect.com/science/article/abs/pii/S0006291X02023331]
- 17. Four translation initiation pathways employed by the ... [https://www.nature.com/articles/srep37905]
- 18. The −10 region adjacent to open reading frames is a ... [https://www.life-science-alliance.org/content/8/11/e202503354]
- 19. AI-directed gene fusing prolongs the evolutionary half-life of ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12487889/]
- 20. Shine–Dalgarno sequence [https://en.wikipedia.org/wiki/Shine%E2%80%93Dalgarno_sequence]
- 21. Shine-Dalgarno Sequence - an overview [https://www.sciencedirect.com/topics/biochemistry-genetics-and-molecular-biology/shine-dalgarno-sequence]
- 22. Correlations between Shine-Dalgarno Sequences and ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC139613/]
- 23. Shine-Dalgarno Sequences Play an Essential Role in the ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC5757275/]
- 24. Propensity for proto-gene emergence in bacteria - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC12542359/]
- 25. Regulation of Leaderless mRNA Translation in Bacteria [https://www.mdpi.com/2076-2607/10/4/723]
- 26. Choreography of human mitochondrial leaderless mRNA ... [https://pubmed.ncbi.nlm.nih.gov/40672251/]
- 27. The Impact of Leadered and Leaderless Gene Structures ... [https://pubmed.ncbi.nlm.nih.gov/32094162/]
- 28. Structural insight into translation initiation of the λcl ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10491246/]
- 29. How Dedicated Ribosomes Translate a Leaderless mRNA [https://www.sciencedirect.com/science/article/abs/pii/S0022283623005405]
- 30. Modeling leaderless transcription and atypical genes results in more... [https://pubmed.ncbi.nlm.nih.gov/29773659/]
- 31. - GeneMarkS 2 [https://exon.gatech.edu/genemarks2.cgi]
- 32. SignalP 6.0 - DTU Health Tech - Bioinformatic Services [https://services.healthtech.dtu.dk/services/SignalP-6.0/]
- 33. SignalP 6.0 predicts all five types of signal peptides using ... [https://www.nature.com/articles/s41587-021-01156-3]
- 34. Signal Peptide Efficiency: From High-Throughput Data to ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC9942255/]
- 35. Signal peptide prediction based on analysis of ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC2286551/]
- 36. SignalP 5.0 - DTU Health Tech - Bioinformatic Services [https://services.healthtech.dtu.dk/services/SignalP-5.0/]
- 37. a comprehensive resource for translation initiation site ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC2238952/]
- 38. Diversity of Translation Initiation Mechanisms across ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC5850609/]
- 39. Target Shuffling [https://www.elderresearch.com/resource/innovations/target-shuffling-process/]
- 40. Design and validation of a reporter mouse to study the ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11262230/]
- 41. Quantitative essentiality in a reduced genome: a functional ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12494982/]
- 42. Automated model-predictive design of synthetic promoters ... [https://www.nature.com/articles/s41467-022-32829-5]
- 43. Integrating transcriptomic and proteomic data for accurate ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC5204337/]
- 44. Integrated Transcriptomic-Proteomic Analysis Using a ... [https://www.sciencedirect.com/science/article/pii/S1535947620337269]
- 45. Hon-yaku: a biology-driven Bayesian methodology for ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC1805508/]
- 46. Ribosome-binding site [https://en.wikipedia.org/wiki/Ribosome-binding_site]
- 47. Comprehensive quantitative modeling of translation efficiency ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10568206/]
- 48. untranslated region in microRNA-mediated gene regulation [https://pmc.ncbi.nlm.nih.gov/articles/PMC4138320/]
- 49. Evidence for the Translation Initiation of Leaderless ... [https://www.sciencedirect.com/science/article/pii/S0021925817477507]
- 50. Influence of 5′-UTR nucleotide composition on translation ... [https://www.sciencedirect.com/science/article/abs/pii/S0923250824001177]
- 51. Translational efficiency is regulated by the length of the 3' ... [https://pubmed.ncbi.nlm.nih.gov/8524291/]
- 52. Enhancing mRNA translation efficiency by introducing ... [https://www.sciencedirect.com/science/article/pii/S2162253125000393]
- 53. Translation efficiency is a determinant of the magnitude ... [https://www.nature.com/articles/s41598-017-13851-w]
- 54. Predicting the translation efficiency of messenger RNA in ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12323635/]
- 55. Leaderless mRNAs in the Spotlight: Ancient but Not ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11633608/]
- 56. Roles for transcript leaders in translation and mRNA decay ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC3668365/]
- 57. Combinatorial optimization of mRNA structure, stability ... [https://www.nature.com/articles/s41467-022-28776-w]
- 58. Enhanced gene regulation by cooperation between mRNA ... [https://www.sciencedirect.com/science/article/pii/S1874939923000056]
- 59. Current Analytical Strategies for mRNA-Based Therapeutics [https://pmc.ncbi.nlm.nih.gov/articles/PMC11990077/]
- 60. mRNA medicine: Recent progresses in chemical ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12311907/]
- 61. Leaderless Transcripts and Small Proteins Are Common ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC4633059/]
- 62. Genome-wide Mapping of Transcriptional Start Sites ... [https://www.sciencedirect.com/science/article/pii/S2211124713006153]
- 63. Translation of a Leaderless Reporter Is Robust During ... [https://www.frontiersin.org/journals/microbiology/articles/10.3389/fmicb.2021.746320/full]
- 64. Mycobacterial RNase E cleaves with a distinct sequence ... [https://www.sciencedirect.com/science/article/pii/S0021925823023402]
- 65. Untranslated region [https://en.wikipedia.org/wiki/Untranslated_region]
- 66. The role of untranslated region variants in Mendelian disease [https://pmc.ncbi.nlm.nih.gov/articles/PMC7617937/]
- 67. regulatory region to bacterial synthetic biology applications [https://pmc.ncbi.nlm.nih.gov/articles/PMC8601185/]
- 68. The quantitative impact of 3'UTRs on gene expression - PubMed [https://pubmed.ncbi.nlm.nih.gov/40586305/]
- 69. 70S-scanning initiation is a novel and frequent ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC4780633/]
- 70. Molecular Mechanism of Scanning and Start Codon Selection ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC3165540/]
- 71. How Dedicated Ribosomes Translate a Leaderless mRNA [https://pmc.ncbi.nlm.nih.gov/articles/PMC11003707/]
- 72. Transcriptome-wide mapping of small ribosomal subunits ... [https://www.nature.com/articles/s42003-025-08804-3]
- 73. attenuation by minocycline enhances longevity and... Translation [https://elifesciences.org/articles/40314]
- 74. of Non-autonomous Mitochondrial Translational ... Regulation Stress [https://pubmed.ncbi.nlm.nih.gov/31340143/]
- 75. Differences in the stability of developmentally regulated ... [https://www.sciencedirect.com/science/article/pii/0012160682902962]
- 76. The stability of an mRNA is influenced by its concentration [https://pmc.ncbi.nlm.nih.gov/articles/PMC5714132/]
- 77. Differential regulation of mRNA stability modulates ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC9936472/]
- 78. UCLA Tool Reveals How mRNA Stability Affects Disease [https://www.technologynetworks.com/proteomics/news/rna-stability-affects-health-risks-404450]
- 79. The utility of protein and mRNA correlation - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC4776753/]
- 80. Conserved cross-domain protein-to-mRNA ratios enable ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12345168/]
- 81. A short translational ramp determines the efficiency of ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC6920384/]
- 82. Translation and mRNA Stability Control - PubMed - NIH [https://pubmed.ncbi.nlm.nih.gov/37001134/]
- 83. Highly stable and efficient templates for mRNA – mRNA ... protein [https://pmc.ncbi.nlm.nih.gov/articles/PMC169963/]
- 84. What is a Model Organism? Moving Beyond E. coli [https://asm.org/articles/2025/february/what-is-a-model-organism-moving-beyond-e-coli]
- 85. Selective Translation of Leaderless mRNAs by Specialized ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC4894548/]
- 86. Ribosomes bind leaderless mRNA in Escherichia coli through ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC2553737/]
- 87. Mycobacterium smegmatis: The Vanguard of Mycobacterial ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC9879119/]